Crystal structures of human 3-hydroxyanthranilate 3,4-dioxygenase with native and non-native metals bound in the active site
- PMID: 28375145
- PMCID: PMC8493610
- DOI: 10.1107/S2059798317002029
Crystal structures of human 3-hydroxyanthranilate 3,4-dioxygenase with native and non-native metals bound in the active site
Abstract
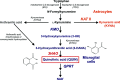
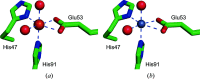
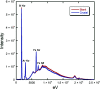
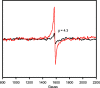
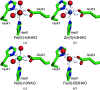
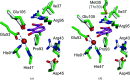
3-Hydroxyanthranilate 3,4-dioxygenase (3HAO) is an enzyme in the microglial branch of the kynurenine pathway of tryptophan degradation. 3HAO is a non-heme iron-containing, ring-cleaving extradiol dioxygenase that catalyzes the addition of both atoms of O2 to the kynurenine pathway metabolite 3-hydroxyanthranilic acid (3-HANA) to form quinolinic acid (QUIN). QUIN is a highly potent excitotoxin that has been implicated in a number of neurodegenerative conditions, making 3HAO a target for pharmacological downregulation. Here, the first crystal structure of human 3HAO with the native iron bound in its active site is presented, together with an additional structure with zinc (a known inhibitor of human 3HAO) bound in the active site. The metal-binding environment is examined both structurally and via inductively coupled plasma mass spectrometry (ICP-MS), X-ray fluorescence spectroscopy (XRF) and electron paramagnetic resonance spectroscopy (EPR). The studies identified Met35 as the source of potential new interactions with substrates and inhibitors, which may prove useful in future therapeutic efforts.
Keywords: 3-hydroxyanthranilate 3,4-dioxygenase; kynurenine pathway; neurodegeneration; oxidoreductase; quinolinic acid.
Figures
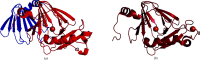

References
-
- Arciero, D. M. & Lipscomb, J. D. (1986). J. Biol. Chem. 261, 2170–2178. - PubMed
-
- Arciero, D. M., Orville, A. M. & Lipscomb, J. D. (1985). J. Biol. Chem. 260, 14035–14044. - PubMed
-
- Bricogne, G., Blanc, E., Brandl, M., Flensburg, C., Keller, P., Paciorek, W., Roversi, P., Sharff, A., Smart, O. S., Vonrhein, C. & Womack, T. O. (2011). BUSTER v.2.10.2. Cambridge: Global Phasing Ltd.
-
- Calderone, V., Trabucco, M., Menin, V., Negro, A. & Zanotti, G. (2002). Biochim. Biophys. Acta, 1596, 283–292. - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

