Modelling the three-dimensional structure of the right-terminal domain of pospiviroids
- PMID: 28386073
- PMCID: PMC5429643
- DOI: 10.1038/s41598-017-00764-x
Modelling the three-dimensional structure of the right-terminal domain of pospiviroids
Abstract
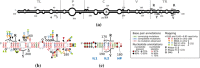
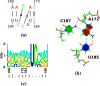
Viroids, the smallest know plant pathogens, consist solely of a circular, single-stranded, non-coding RNA. Thus for all of their biological functions, like replication, processing, and transport, they have to present sequence or structural features to exploit host proteins. Viroid binding protein 1 (Virp1) is indispensable for replication of pospiviroids, the largest genus of viroids, in a host plant as well as in protoplasts. Virp1 is known to bind at two sites in the terminal right (TR) domain of pospiviroids; each site consists of a purine- (R-) and a pyrimidine- (Y-)rich motif that are partially base-paired to each other. Here we model the important structural features of the domain and show that it contains an internal loop of two Y · Y cis Watson-Crick/Watson-Crick (cWW) pairs, an asymmetric internal loop including a cWW and a trans Watson/Hoogsteen pair, and a thermodynamically quite stable hairpin loop with several stacking interactions. These features are discussed in connection to the known biological functions of the TR domain.
Conflict of interest statement
The authors declare that they have no competing interests.
Figures





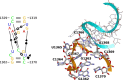
Similar articles
-
Dissecting the secondary structure of the circular RNA of a nuclear viroid in vivo: A "naked" rod-like conformation similar but not identical to that observed in vitro.RNA Biol. 2017 Aug 3;14(8):1046-1054. doi: 10.1080/15476286.2016.1223005. Epub 2016 Aug 30. RNA Biol. 2017. PMID: 27574720 Free PMC article.
-
In silico survey of the central conserved regions in viroids of the Pospiviroidae family for conserved asymmetric loop structures.RNA. 2019 Aug;25(8):985-1003. doi: 10.1261/rna.070409.119. Epub 2019 May 23. RNA. 2019. PMID: 31123078 Free PMC article.
-
Structural differences within the loop E motif imply alternative mechanisms of viroid processing.RNA. 2007 Jun;13(6):824-34. doi: 10.1261/rna.452307. Epub 2007 Apr 16. RNA. 2007. PMID: 17438124 Free PMC article.
-
Structure and Associated Biological Functions of Viroids.Adv Virus Res. 2016;94:141-72. doi: 10.1016/bs.aivir.2015.11.002. Epub 2016 Mar 2. Adv Virus Res. 2016. PMID: 26997592 Review.
-
Viroid: a useful model for studying the basic principles of infection and RNA biology.Mol Plant Microbe Interact. 2007 Jan;20(1):7-20. doi: 10.1094/MPMI-20-0007. Mol Plant Microbe Interact. 2007. PMID: 17249418 Review.
Cited by
-
Predicting the Structure of a Viroid : Structure, Structure Distribution, Consensus Structure, and Structure Drawing.Methods Mol Biol. 2022;2316:331-371. doi: 10.1007/978-1-0716-1464-8_26. Methods Mol Biol. 2022. PMID: 34845705
-
Derailing the host machinery to achieve replication: how viroid and viroid-like RNAs successfully copy their genomes in hostile territory.RNA Biol. 2025 Dec;22(1):1-19. doi: 10.1080/15476286.2025.2538269. Epub 2025 Aug 26. RNA Biol. 2025. PMID: 40720211 Free PMC article. Review.
-
A nuclear import pathway exploited by pathogenic noncoding RNAs.Plant Cell. 2022 Sep 27;34(10):3543-3556. doi: 10.1093/plcell/koac210. Plant Cell. 2022. PMID: 35877068 Free PMC article.
-
Viroid research and its significance for RNA technology and basic biochemistry.Nucleic Acids Res. 2018 Nov 16;46(20):10563-10576. doi: 10.1093/nar/gky903. Nucleic Acids Res. 2018. PMID: 30304486 Free PMC article. Review.
-
Whole-Genome Deep Sequencing Reveals Host-Driven in-planta Evolution of Columnea Latent Viroid (CLVd) Quasi-Species Populations.Int J Mol Sci. 2020 May 5;21(9):3262. doi: 10.3390/ijms21093262. Int J Mol Sci. 2020. PMID: 32380694 Free PMC article.
References
-
- Schindler I-M, Mühlbach H-P. Involvement of nuclear DNA-dependent RNA polymerases in potato spindle tuber viroid replication: a reevaluation. Plant Sci. 1992;84:221–229. doi: 10.1016/0168-9452(92)90138-C. - DOI
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials

