Activation-induced deoxycytidine deaminase: Structural basis for favoring WRC hot motif specificities unique among APOBEC family members
- PMID: 28388461
- PMCID: PMC6408873
- DOI: 10.1016/j.dnarep.2017.03.007
Activation-induced deoxycytidine deaminase: Structural basis for favoring WRC hot motif specificities unique among APOBEC family members
Keywords: AID X-ray crystal structure; Antibody diversity; C-deamination motif selectivity; IgV somatic hypermutation.
Figures
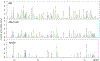


References
-
- Pham P, Bransteitter R, Petruska J, Goodman MF, Processive AID-catalysed cytosine deamination on single-stranded DNA simulates somatic hypermutation, Nature, 424 (2003) 103–107. - PubMed
-
- Beale RC, Petersen-Mahrt SK, Watt IN, Harris RS, Rada C, Neuberger MS, Comparison of the differential context-dependence of DNA deamination by APOBEC enzymes: correlation with mutation spectra in vivo, J Mol Biol, 337 (2004) 585–596. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

