A comprehensive hybridization model allows whole HERV transcriptome profiling using high density microarray
- PMID: 28390408
- PMCID: PMC5385096
- DOI: 10.1186/s12864-017-3669-7
A comprehensive hybridization model allows whole HERV transcriptome profiling using high density microarray
Abstract
Background: Human endogenous retroviruses (HERVs) have received much attention for their implications in the etiology of many human diseases and their profound effect on evolution. Notably, recent studies have highlighted associations between HERVs expression and cancers (Yu et al., Int J Mol Med 32, 2013), autoimmunity (Balada et al., Int Rev Immunol 29:351-370, 2010) and neurological (Christensen, J Neuroimmune Pharmacol 5:326-335, 2010) conditions. Their repetitive nature makes their study particularly challenging, where expression studies have largely focused on individual loci (De Parseval et al., J Virol 77:10414-10422, 2003) or general trends within families (Forsman et al., J Virol Methods 129:16-30, 2005; Seifarth et al., J Virol 79:341-352, 2005; Pichon et al., Nucleic Acids Res 34:e46, 2006).
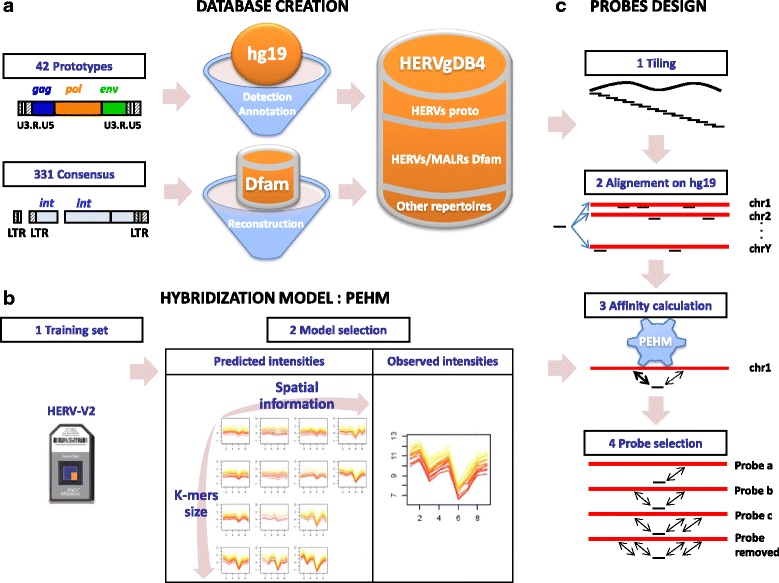
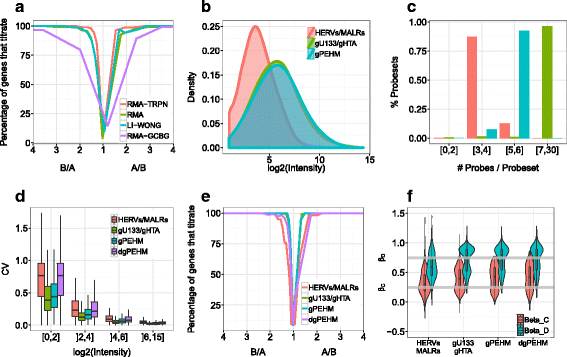
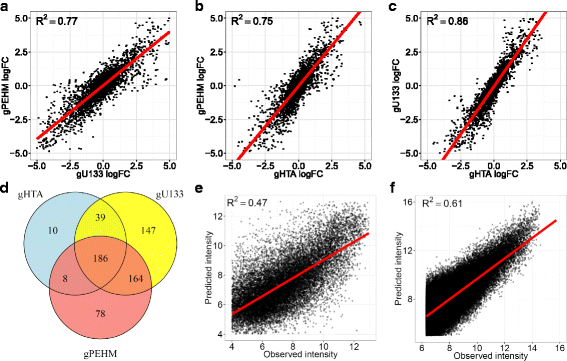
Methods: To refine our understanding of HERVs activity, we introduce here a new microarray, HERV-V3. This work was made possible by the careful detection and annotation of genomic HERV/MaLR sequences as well as the development of a new hybridization model, allowing the optimization of probe performances and the control of cross-reactions. RESULTS: HERV-V3 offers an almost complete coverage of HERVs and their ancestors (mammalian apparent LTR-retrotransposons, MaLRs) at the locus level along with four other repertoires (active LINE-1 elements, lncRNA, a selection of 1559 human genes and common infectious viruses). We demonstrate that HERV-V3 analytical performances are comparable with commercial Affymetrix arrays, and that for a selection of tissue/pathological specific loci, the patterns of expression measured on HERV-V3 is consistent with those reported in the literature.
Conclusions: Given its large HERVs/MaLRs coverage and additional repertoires, HERV-V3 opens the door to multiple applications such as enhancers and alternative promoters identification, biomarkers identification as well as the characterization of genes and HERVs/MaLRs modulation caused by viral infection.
Keywords: Biostatistics; Microarray; Repetitive elements; Transcriptomics.
Figures

References
-
- Lander ES, Linton LM, Birren B, Nusbaum C, Zody MC, Baldwin J, Devon K, Dewar K, Doyle M, FitzHugh W, Evans GA, Athanasiou M, Schultz R, Patrinos A, Morgan MJ. Initial sequencing and analysis of the human genome. Nature. 2001;409(6822):860–921. - PubMed
-
- Mager DL, Medstrand P. Retroviral repeat sequences. Chichester: eLS. Wiley; 2005.
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources
Other Literature Sources

