Accurate Classification of Protein Subcellular Localization from High-Throughput Microscopy Images Using Deep Learning
- PMID: 28391243
- PMCID: PMC5427497
- DOI: 10.1534/g3.116.033654
Accurate Classification of Protein Subcellular Localization from High-Throughput Microscopy Images Using Deep Learning
Abstract
High-throughput microscopy of many single cells generates high-dimensional data that are far from straightforward to analyze. One important problem is automatically detecting the cellular compartment where a fluorescently-tagged protein resides, a task relatively simple for an experienced human, but difficult to automate on a computer. Here, we train an 11-layer neural network on data from mapping thousands of yeast proteins, achieving per cell localization classification accuracy of 91%, and per protein accuracy of 99% on held-out images. We confirm that low-level network features correspond to basic image characteristics, while deeper layers separate localization classes. Using this network as a feature calculator, we train standard classifiers that assign proteins to previously unseen compartments after observing only a small number of training examples. Our results are the most accurate subcellular localization classifications to date, and demonstrate the usefulness of deep learning for high-throughput microscopy.
Keywords: deep learning; high-content screening; machine learning; microscopy; yeast.
Copyright © 2017 Parnamaa and Parts.
Figures

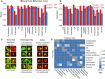


Similar articles
-
An Automatic Segmentation Method Combining an Active Contour Model and a Classification Technique for Detecting Polycomb-group Proteinsin High-Throughput Microscopy Images.Methods Mol Biol. 2016;1480:181-97. doi: 10.1007/978-1-4939-6380-5_16. Methods Mol Biol. 2016. PMID: 27659985
-
A multiresolution approach to automated classification of protein subcellular location images.BMC Bioinformatics. 2007 Jun 19;8:210. doi: 10.1186/1471-2105-8-210. BMC Bioinformatics. 2007. PMID: 17578580 Free PMC article.
-
Automated analysis of high-content microscopy data with deep learning.Mol Syst Biol. 2017 Apr 18;13(4):924. doi: 10.15252/msb.20177551. Mol Syst Biol. 2017. PMID: 28420678 Free PMC article.
-
Machine learning and computer vision approaches for phenotypic profiling.J Cell Biol. 2017 Jan 2;216(1):65-71. doi: 10.1083/jcb.201610026. Epub 2016 Dec 9. J Cell Biol. 2017. PMID: 27940887 Free PMC article. Review.
-
Computer vision for high content screening.Crit Rev Biochem Mol Biol. 2016;51(2):102-9. doi: 10.3109/10409238.2015.1135868. Epub 2016 Jan 24. Crit Rev Biochem Mol Biol. 2016. PMID: 26806341 Review.
Cited by
-
HAR_Locator: a novel protein subcellular location prediction model of immunohistochemistry images based on hybrid attention modules and residual units.Front Mol Biosci. 2023 Aug 17;10:1171429. doi: 10.3389/fmolb.2023.1171429. eCollection 2023. Front Mol Biosci. 2023. PMID: 37664182 Free PMC article.
-
Self-Learning Microfluidic Platform for Single-Cell Imaging and Classification in Flow.Micromachines (Basel). 2019 May 9;10(5):311. doi: 10.3390/mi10050311. Micromachines (Basel). 2019. PMID: 31075890 Free PMC article.
-
Deep learning unlocks label-free viability assessment of cancer spheroids in microfluidics.Lab Chip. 2024 Jun 11;24(12):3169-3182. doi: 10.1039/d4lc00197d. Lab Chip. 2024. PMID: 38804084 Free PMC article.
-
Single-cell image analysis to explore cell-to-cell heterogeneity in isogenic populations.Cell Syst. 2021 Jun 16;12(6):608-621. doi: 10.1016/j.cels.2021.05.010. Cell Syst. 2021. PMID: 34139168 Free PMC article. Review.
-
Transfer learning for versatile and training free high content screening analyses.Sci Rep. 2023 Dec 18;13(1):22599. doi: 10.1038/s41598-023-49554-8. Sci Rep. 2023. PMID: 38114550 Free PMC article.
References
-
- Alipanahi B., Delong A., Weirauch M. T., Frey B. J., 2015. Predicting the sequence specificities of DNA- and RNA-binding proteins by deep learning. Nat. Biotechnol. 33: 831–838. - PubMed
-
- Boland M. V., Murphy R. F., 2001. A neural network classifier capable of recognizing the patterns of all major subcellular structures in fluorescence microscope images of HeLa cells. Bioinformatics 17(12): 1213–1223. - PubMed
-
- Boland M. V., Markey M. K., Murphy R. F., 1998. Automated recognition of patterns characteristic of subcellular structures in fluorescence microscopy images. Cytometry 33(3): 366–375. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
