Mechanistic Analysis of Fluorescence Quenching of Reduced Nicotinamide Adenine Dinucleotide by Oxamate in Lactate Dehydrogenase Ternary Complexes
- PMID: 28391608
- PMCID: PMC5603363
- DOI: 10.1111/php.12775
Mechanistic Analysis of Fluorescence Quenching of Reduced Nicotinamide Adenine Dinucleotide by Oxamate in Lactate Dehydrogenase Ternary Complexes
Abstract
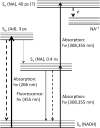
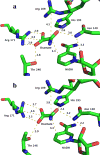
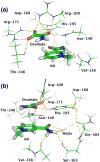
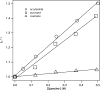
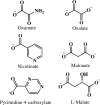
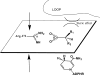
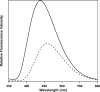
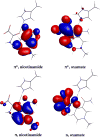
Fluorescence of Reduced Nicotinamide Adenine Dinucleotide (NADH) is extensively employed in studies of oxidoreductases. A substantial amount of static and kinetic work has focused on the binding of pyruvate or substrate mimic oxamate to the binary complex of lactate dehydrogenase (LDH)-NADH where substantial fluorescence quenching is typically observed. However, the quenching mechanism is not well understood limiting structural interpretation. Based on time-dependent density functional theory (TDDFT) computations with cam-B3LYP functional in conjunction with the analysis of previous experimental results, we propose that bound oxamate acts as an electron acceptor in the quenching of fluorescence of NADH in the ternary complex, where a charge transfer (CT) state characterized by excitation from the highest occupied molecular orbital (HOMO) of the nicotinamide moiety of NADH to the lowest unoccupied molecular orbital (LUMO) of oxamate exists close to the locally excited (LE) state involving only the nicotinamide moiety. Efficient quenching in the encounter complex like in pig heart LDH requires that oxamate forms a salt bridge with Arg-171 and hydrogen bonds with His-195, Thr-246 and Asn-140. Further structural rearrangement and loop closure, which also brings about another hydrogen bond between oxamate and Arg-109, will increase the rate of fluorescence quenching as well.
© 2017 The American Society of Photobiology.
Figures

Similar articles
-
Mechanism for Fluorescence Quenching of Tryptophan by Oxamate and Pyruvate: Conjugation and Solvation-Induced Photoinduced Electron Transfer.J Phys Chem B. 2018 Jun 28;122(25):6483-6490. doi: 10.1021/acs.jpcb.8b02433. Epub 2018 Jun 15. J Phys Chem B. 2018. PMID: 29860828 Free PMC article.
-
Kinetic mechanism of the endogenous lactate dehydrogenase activity of duck epsilon-crystallin.Arch Biochem Biophys. 1991 Feb 1;284(2):285-91. doi: 10.1016/0003-9861(91)90297-v. Arch Biochem Biophys. 1991. PMID: 1989512
-
Equilibrium isotope effect on ternary complex formation of [1-18O]oxamate with NADH and lactate dehydrogenase.Biochemistry. 1995 May 9;34(18):6050-8. doi: 10.1021/bi00018a007. Biochemistry. 1995. PMID: 7742308
-
The use of ternary complexes to study ionizations and isomerizations during catalysis by lactate dehydrogenase.Biochem J. 1973 Apr;131(4):739-48. doi: 10.1042/bj1310739. Biochem J. 1973. PMID: 4352914 Free PMC article.
-
Slow structural changes shown by the 3-nitrotyrosine-237 residue in pig heart [Tyr(3NO2)237] lactate dehydrogenase.Biochem J. 1982 Mar 1;201(3):465-71. doi: 10.1042/bj2010465. Biochem J. 1982. PMID: 7092806 Free PMC article.
Cited by
-
Mechanism for Fluorescence Quenching of Tryptophan by Oxamate and Pyruvate: Conjugation and Solvation-Induced Photoinduced Electron Transfer.J Phys Chem B. 2018 Jun 28;122(25):6483-6490. doi: 10.1021/acs.jpcb.8b02433. Epub 2018 Jun 15. J Phys Chem B. 2018. PMID: 29860828 Free PMC article.
-
A short guide on blue fluorescent proteins: limits and perspectives.Appl Microbiol Biotechnol. 2024 Feb 14;108(1):208. doi: 10.1007/s00253-024-13012-w. Appl Microbiol Biotechnol. 2024. PMID: 38353763 Free PMC article. Review.
-
Resolution of Submillisecond Kinetics of Multiple Reaction Pathways for Lactate Dehydrogenase.Biophys J. 2017 May 9;112(9):1852-1862. doi: 10.1016/j.bpj.2017.03.031. Biophys J. 2017. PMID: 28494956 Free PMC article.
-
NAD(P)H binding configurations revealed by time-resolved fluorescence and two-photon absorption.Biophys J. 2023 Apr 4;122(7):1240-1253. doi: 10.1016/j.bpj.2023.02.014. Epub 2023 Feb 14. Biophys J. 2023. PMID: 36793214 Free PMC article.
-
Small molecule cores demonstrate non-competitive inhibition of lactate dehydrogenase.Medchemcomm. 2018 Jul 13;9(8):1369-1376. doi: 10.1039/c8md00309b. eCollection 2018 Aug 1. Medchemcomm. 2018. PMID: 30151092 Free PMC article.
References
-
- Wu Y, Zheng W, Qu JY. Sensing cell metabolism by time-resolved autofluorescence. Opt Lett. 2006;31:3122. - PubMed
-
- Wang H-W, Wei Y-H, Guo H-W. Reduced Nicotinamide Adenine Dinucleotide (NADH) Fluorescence for the Detection of Cell Death. Anticancer Agents Med Chem. 2009;9:1012–1017. - PubMed
-
- Bartolome F, Abramov AY. Measurement of mitochondrial NADH and FAD autofluorescence in live cells. In: Weissig V, Edeas M, editors. Mitochondrial Medicine: Volume I, Probing Mitochondrial function 1264. Springer; New York: 2015. pp. 263–270. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

