Neurogenetic analysis of childhood disintegrative disorder
- PMID: 28392909
- PMCID: PMC5379515
- DOI: 10.1186/s13229-017-0133-0
Neurogenetic analysis of childhood disintegrative disorder
Abstract
Background: Childhood disintegrative disorder (CDD) is a rare form of autism spectrum disorder (ASD) of unknown etiology. It is characterized by late-onset regression leading to significant intellectual disability (ID) and severe autism. Although there are phenotypic differences between CDD and other forms of ASD, it is unclear if there are neurobiological differences.
Methods: We pursued a multidisciplinary study of CDD (n = 17) and three comparison groups: low-functioning ASD (n = 12), high-functioning ASD (n = 50), and typically developing (n = 26) individuals. We performed whole-exome sequencing (WES), copy number variant (CNV), and gene expression analyses of CDD and, on subsets of each cohort, non-sedated functional magnetic resonance imaging (fMRI) while viewing socioemotional (faces) and non-socioemotional (houses) stimuli and eye tracking while viewing emotional faces.
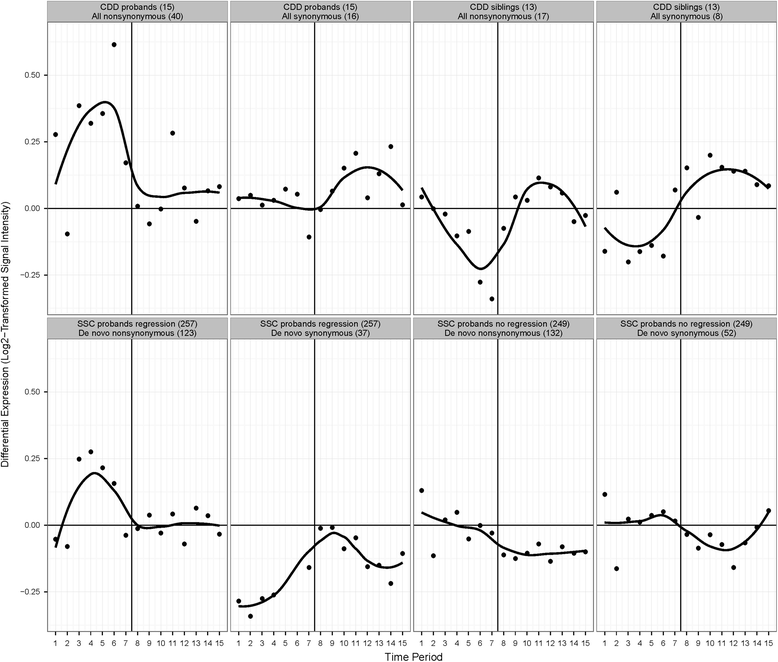
Results: We observed potential differences between CDD and other forms of ASD. WES and CNV analyses identified one or more rare de novo, homozygous, and/or hemizygous (mother-to-son transmission on chrX) variants for most probands that were not shared by unaffected sibling controls. There were no clearly deleterious variants or highly recurrent candidate genes. Candidate genes that were found to be most conserved at variant position and most intolerant of variation, such as TRRAP, ZNF236, and KIAA2018, play a role or may be involved in transcription. Using the human BrainSpan transcriptome dataset, CDD candidate genes were found to be more highly expressed in non-neocortical regions than neocortical regions. This expression profile was similar to that of an independent cohort of ASD probands with regression. The non-neocortical regions overlapped with those identified by fMRI as abnormally hyperactive in response to viewing faces, such as the thalamus, cerebellum, caudate, and hippocampus. Eye-tracking analysis showed that, among individuals with ASD, subjects with CDD focused on eyes the most when shown pictures of faces.
Conclusions: Given that cohort sizes were limited by the rarity of CDD, and the challenges of conducting non-sedated fMRI and eye tracking in subjects with ASD and significant ID, this is an exploratory study designed to investigate the neurobiological features of CDD. In addition to reporting the first multimodal analysis of CDD, a combination of fMRI and eye-tracking analyses are being presented for the first time for low-functioning individuals with ASD. Our results suggest differences between CDD and other forms of ASD on the neurobiological as well as clinical level.
Keywords: Autism spectrum disorder (ASD); Childhood disintegrative disorder (CDD); Eye tracking; Functional magnetic resonance imaging (fMRI); Genetics; Intellectual disability (ID); Regression.
Figures





References
-
- American Psychiatric Association . Diagnostic and Statistical Manual of Mental Disorders: DSM-5. 5. Washington: American Psychiatric Association; 2013. Autism spectrum disorder; pp. 50–9.
-
- Kanner L. Autistic disturbances of affective contact. Nerv Child. 1943;2:217–50. - PubMed
-
- Heller T. Dementia infantilis. Zeitschrift fur die Erforschung und Behandlung des Jugenlichen, Schwachsinns. 1908;2:17–28.
-
- World Health Organization . The ICD-10 classification of mental and behavioural disorders: clinical descriptions and diagnostic guidelines. Geneva: World Health Organization; 1992.
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases

