Genome-to-genome analysis highlights the effect of the human innate and adaptive immune systems on the hepatitis C virus
- PMID: 28394351
- PMCID: PMC5873514
- DOI: 10.1038/ng.3835
Genome-to-genome analysis highlights the effect of the human innate and adaptive immune systems on the hepatitis C virus
Abstract
Outcomes of hepatitis C virus (HCV) infection and treatment depend on viral and host genetic factors. Here we use human genome-wide genotyping arrays and new whole-genome HCV viral sequencing technologies to perform a systematic genome-to-genome study of 542 individuals who were chronically infected with HCV, predominantly genotype 3. We show that both alleles of genes encoding human leukocyte antigen molecules and genes encoding components of the interferon lambda innate immune system drive viral polymorphism. Additionally, we show that IFNL4 genotypes determine HCV viral load through a mechanism dependent on a specific amino acid residue in the HCV NS5A protein. These findings highlight the interplay between the innate immune system and the viral genome in HCV control.
Conflict of interest statement
The authors disclose the following: G.R.F: Grants Consulting and Speaker/Advisory Board: AbbVie, Alcura, Bristol-Myers Squibb, Gilead, Janssen, GlaxoSmithKline, Merck, Roche, Springbank, Idenix, Tekmira, Novartis. G.M. is a partner in Peptide Groove LLP, which commercializes HLA*IMP.
Figures
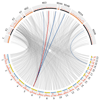
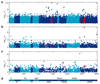
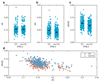
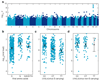

References
-
- Mohd Hanafiah K, Groeger J, Flaxman AD, Wiersma ST. Global epidemiology of hepatitis C virus infection: new estimates of age-specific antibody to HCV seroprevalence. Hepatology. 2013;57:1333–1342. - PubMed
-
- Ge D, et al. Genetic variation in IL28B predicts hepatitis C treatment-induced viral clearance. Nature. 2009;461:399–401. - PubMed
-
- Rauch A, et al. Genetic variation in IL28B is associated with chronic hepatitis C and treatment failure: a genome-wide association study. Gastroenterology. 2010;138:1338–45. 1345–7. - PubMed
-
- Suppiah V, et al. IL28B is associated with response to chronic hepatitis C interferon-alpha and ribavirin therapy. Nat Genet. 2009;41:1100–1104. - PubMed
MeSH terms
Substances
Grants and funding
- MC_UU_12014/1/MRC_/Medical Research Council/United Kingdom
- C0365/MRF_/MRF_/United Kingdom
- EME/14/02/17/DH_/Department of Health/United Kingdom
- G1100247/MRC_/Medical Research Council/United Kingdom
- U19 AI082630/AI/NIAID NIH HHS/United States
- MR/P025064/1/MRC_/Medical Research Council/United Kingdom
- 109965/WT_/Wellcome Trust/United Kingdom
- G0400802/MRC_/Medical Research Council/United Kingdom
- MR/K01532X/1/MRC_/Medical Research Council/United Kingdom
- MC_EX_UU_G1000717/MRC_/Medical Research Council/United Kingdom
- 109965/Z/15/Z/WT_/Wellcome Trust/United Kingdom
- RP-2016-07-012/DH_/Department of Health/United Kingdom
- G0801976/MRC_/Medical Research Council/United Kingdom
- MC_UP_A390_1107/MRC_/Medical Research Council/United Kingdom
- G1001738/MRC_/Medical Research Council/United Kingdom
LinkOut - more resources
Full Text Sources
Other Literature Sources

