Adaptation and evolution of deep-sea scale worms (Annelida: Polynoidae): insights from transcriptome comparison with a shallow-water species
- PMID: 28397791
- PMCID: PMC5387418
- DOI: 10.1038/srep46205
Adaptation and evolution of deep-sea scale worms (Annelida: Polynoidae): insights from transcriptome comparison with a shallow-water species
Abstract
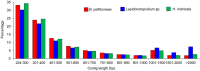
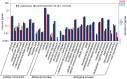
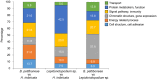
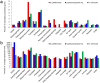
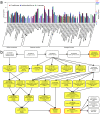
Polynoid scale worms (Polynoidae, Annelida) invaded deep-sea chemosynthesis-based ecosystems approximately 60 million years ago, but little is known about their genetic adaptation to the extreme deep-sea environment. In this study, we reported the first two transcriptomes of deep-sea polynoids (Branchipolynoe pettiboneae, Lepidonotopodium sp.) and compared them with the transcriptome of a shallow-water polynoid (Harmothoe imbricata). We determined codon and amino acid usage, positive selected genes, highly expressed genes and putative duplicated genes. Transcriptome assembly produced 98,806 to 225,709 contigs in the three species. There were more positively charged amino acids (i.e., histidine and arginine) and less negatively charged amino acids (i.e., aspartic acid and glutamic acid) in the deep-sea species. There were 120 genes showing clear evidence of positive selection. Among the 10% most highly expressed genes, there were more hemoglobin genes with high expression levels in both deep-sea species. The duplicated genes related to DNA recombination and metabolism, and gene expression were only enriched in deep-sea species. Deep-sea scale worms adopted two strategies of adaptation to hypoxia in the chemosynthesis-based habitats (i.e., rapid evolution of tetra-domain hemoglobin in Branchipolynoe or high expression of single-domain hemoglobin in Lepidonotopodium sp.).
Conflict of interest statement
The authors declare no competing financial interests.
Figures


References
-
- Levin L. A. Deep-ocean life where oxygen is scarce. Am. Sci. 90, 436–444 (2002).
-
- Levin L. A. et al.. Hydrothermal Vents and Methane Seeps: Rethinking the Sphere of Influence. Front. Mar. Sci, 3, 72 (2016).
-
- Watanabe H., Fujikura K., Kojima S., Miyazaki J. I. & Fujiwara Y. Japan: vents and seeps in close proximity. In The Vent and Seep Biota (ed. Kiel S.) 379–401 (Springer Netherlands, 2010).
-
- Minic Z., Serre V. & Hervé G. Adaptation of organisms to extreme conditions of deep-sea hydrothermal vents. C. R. Biol. 329, 527–540 (2006). - PubMed
-
- McMullin E. R., Bergquist D. C. & Fisher C. R. Metazoans in extreme environments: adaptations of hydrothermal vent and hydrocarbon seep fauna. Gravitational Space Res. 13, 13–24 (2007). - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

