Defining the transcriptomic landscape of the developing enteric nervous system and its cellular environment
- PMID: 28403821
- PMCID: PMC5389105
- DOI: 10.1186/s12864-017-3653-2
Defining the transcriptomic landscape of the developing enteric nervous system and its cellular environment
Abstract
Background: Motility and the coordination of moving food through the gastrointestinal tract rely on a complex network of neurons known as the enteric nervous system (ENS). Despite its critical function, many of the molecular mechanisms that direct the development of the ENS and the elaboration of neural network connections remain unknown. The goal of this study was to transcriptionally identify molecular pathways and candidate genes that drive specification, differentiation and the neural circuitry of specific neural progenitors, the phox2b expressing ENS cell lineage, during normal enteric nervous system development. Because ENS development is tightly linked to its environment, the transcriptional landscape of the cellular environment of the intestine was also analyzed.
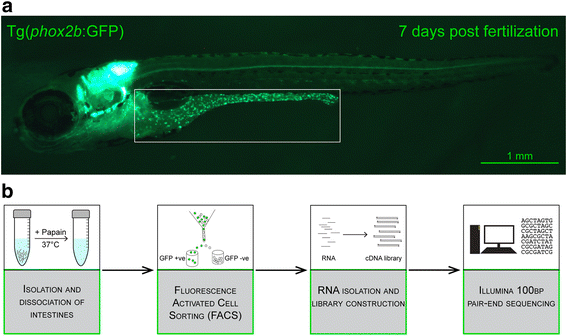
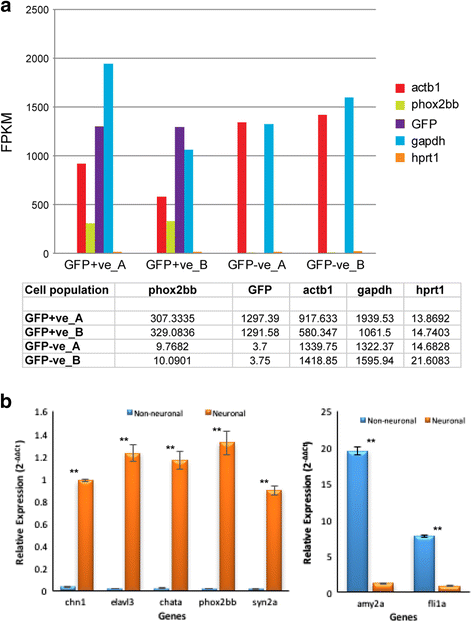
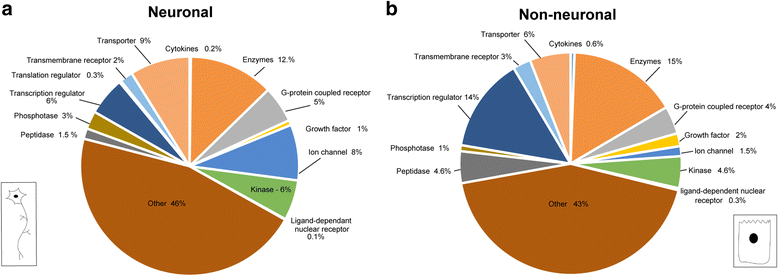
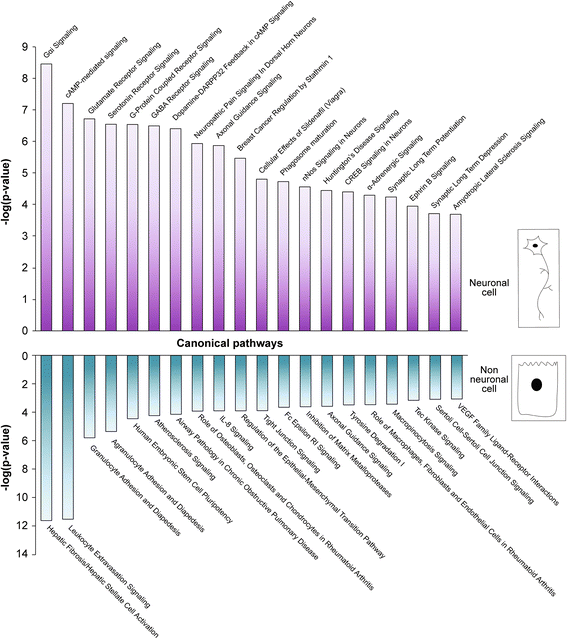
Results: Thousands of zebrafish intestines were manually dissected from a transgenic line expressing green fluorescent protein under the phox2b regulatory elements [Tg(phox2b:EGFP) w37 ]. Fluorescence-activated cell sorting was used to separate GFP-positive phox2b expressing ENS progenitor and derivatives from GFP-negative intestinal cells. RNA-seq was performed to obtain accurate, reproducible transcriptional profiles and the unbiased detection of low level transcripts. Analysis revealed genes and pathways that may function in ENS cell determination, genes that may be identifiers of different ENS subtypes, and genes that define the non-neural cellular microenvironment of the ENS. Differential expression analysis between the two cell populations revealed the expected neuronal nature of the phox2b expressing lineage including the enrichment for genes required for neurogenesis and synaptogenesis, and identified many novel genes not previously associated with ENS development. Pathway analysis pointed to a high level of G-protein coupled pathway activation, and identified novel roles for candidate pathways such as the Nogo/Reticulon axon guidance pathway in ENS development.
Conclusion: We report the comprehensive gene expression profiles of a lineage-specific population of enteric progenitors, their derivatives, and their microenvironment during normal enteric nervous system development. Our results confirm previously implicated genes and pathways required for ENS development, and also identify scores of novel candidate genes and pathways. Thus, our dataset suggests various potential mechanisms that drive ENS development facilitating characterization and discovery of novel therapeutic strategies to improve gastrointestinal disorders.
Keywords: Enteric nervous system; Hirschsprungs; Neural crest; RNA-sequencing; Transcriptome; Zebrafish; phox2b.
Figures


Similar articles
-
Transcription and Signaling Regulators in Developing Neuronal Subtypes of Mouse and Human Enteric Nervous System.Gastroenterology. 2018 Feb;154(3):624-636. doi: 10.1053/j.gastro.2017.10.005. Epub 2017 Oct 12. Gastroenterology. 2018. PMID: 29031500 Free PMC article.
-
Neuronal Differentiation in Schwann Cell Lineage Underlies Postnatal Neurogenesis in the Enteric Nervous System.J Neurosci. 2015 Jul 8;35(27):9879-88. doi: 10.1523/JNEUROSCI.1239-15.2015. J Neurosci. 2015. PMID: 26156989 Free PMC article.
-
A Rapid F0 CRISPR Screen in Zebrafish to Identify Regulator Genes of Neuronal Development in the Enteric Nervous System.Neurogastroenterol Motil. 2025 May;37(5):e70009. doi: 10.1111/nmo.70009. Epub 2025 Apr 6. Neurogastroenterol Motil. 2025. PMID: 40189908 Free PMC article.
-
Gut feelings: Studying enteric nervous system development, function, and disease in the zebrafish model system.Dev Dyn. 2018 Feb;247(2):268-278. doi: 10.1002/dvdy.24597. Epub 2017 Oct 23. Dev Dyn. 2018. PMID: 28975691 Review.
-
Enteric nervous system development: migration, differentiation, and disease.Am J Physiol Gastrointest Liver Physiol. 2013 Jul 1;305(1):G1-24. doi: 10.1152/ajpgi.00452.2012. Epub 2013 May 2. Am J Physiol Gastrointest Liver Physiol. 2013. PMID: 23639815 Free PMC article. Review.
Cited by
-
Regulatory Logic Underlying Diversification of the Neural Crest.Trends Genet. 2017 Oct;33(10):715-727. doi: 10.1016/j.tig.2017.07.015. Epub 2017 Aug 26. Trends Genet. 2017. PMID: 28851604 Free PMC article. Review.
-
Mouse intestinal tuft cells express advillin but not villin.Sci Rep. 2020 Jun 1;10(1):8877. doi: 10.1038/s41598-020-65469-0. Sci Rep. 2020. PMID: 32483224 Free PMC article.
-
Development, Diversity, and Neurogenic Capacity of Enteric Glia.Front Cell Dev Biol. 2022 Jan 17;9:775102. doi: 10.3389/fcell.2021.775102. eCollection 2021. Front Cell Dev Biol. 2022. PMID: 35111752 Free PMC article. Review.
-
Unbiased characterization of the larval zebrafish enteric nervous system at a single cell transcriptomic level.iScience. 2023 Jun 8;26(7):107070. doi: 10.1016/j.isci.2023.107070. eCollection 2023 Jul 21. iScience. 2023. PMID: 37426341 Free PMC article.
-
Identification of Genetic Susceptibility Factors Associated with Canine Gastric Dilatation-Volvulus.Genes (Basel). 2020 Nov 5;11(11):1313. doi: 10.3390/genes11111313. Genes (Basel). 2020. PMID: 33167491 Free PMC article.
References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Miscellaneous

