Loss of Arf4 causes severe degeneration of the exocrine pancreas but not cystic kidney disease or retinal degeneration
- PMID: 28410364
- PMCID: PMC5409180
- DOI: 10.1371/journal.pgen.1006740
Loss of Arf4 causes severe degeneration of the exocrine pancreas but not cystic kidney disease or retinal degeneration
Abstract
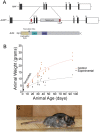
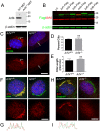
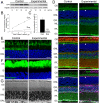
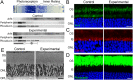
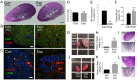
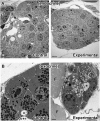
Arf4 is proposed to be a critical regulator of membrane protein trafficking in early secretory pathway. More recently, Arf4 was also implicated in regulating ciliary trafficking, however, this has not been comprehensively tested in vivo. To directly address Arf4's role in ciliary transport, we deleted Arf4 specifically in either rod photoreceptor cells, kidney, or globally during the early postnatal period. Arf4 deletion in photoreceptors did not cause protein mislocalization or retinal degeneration, as expected if Arf4 played a role in protein transport to the ciliary outer segment. Likewise, Arf4 deletion in kidney did not cause cystic disease, as expected if Arf4 were involved in general ciliary trafficking. In contrast, global Arf4 deletion in the early postnatal period resulted in growth restriction, severe pancreatic degeneration and early death. These findings are consistent with Arf4 playing a critical role in endomembrane trafficking, particularly in the pancreas, but not in ciliary function.
Conflict of interest statement
The authors have declared that no competing interests exist.
Figures


Similar articles
-
The ins and outs of the Arf4-based ciliary membrane-targeting complex.Small GTPases. 2021 Jan;12(1):1-12. doi: 10.1080/21541248.2019.1616355. Epub 2019 May 17. Small GTPases. 2021. PMID: 31068062 Free PMC article. Review.
-
Arf4 is required for Mammalian development but dispensable for ciliary assembly.PLoS Genet. 2014 Feb 20;10(2):e1004170. doi: 10.1371/journal.pgen.1004170. eCollection 2014 Feb. PLoS Genet. 2014. PMID: 24586199 Free PMC article.
-
Ciliary targeting motif VxPx directs assembly of a trafficking module through Arf4.EMBO J. 2009 Feb 4;28(3):183-92. doi: 10.1038/emboj.2008.267. Epub 2009 Jan 15. EMBO J. 2009. PMID: 19153612 Free PMC article.
-
Cholesterol 25-Hydroxylase Protects Against Diabetic Kidney Disease by Regulating ADP Ribosylation Factor 4.Adv Sci (Weinh). 2024 Aug;11(29):e2309642. doi: 10.1002/advs.202309642. Epub 2024 May 30. Adv Sci (Weinh). 2024. PMID: 38816950 Free PMC article.
-
Non-syndromic retinal ciliopathies: translating gene discovery into therapy.Hum Mol Genet. 2012 Oct 15;21(R1):R111-24. doi: 10.1093/hmg/dds298. Epub 2012 Jul 26. Hum Mol Genet. 2012. PMID: 22843501 Review.
Cited by
-
The ins and outs of the Arf4-based ciliary membrane-targeting complex.Small GTPases. 2021 Jan;12(1):1-12. doi: 10.1080/21541248.2019.1616355. Epub 2019 May 17. Small GTPases. 2021. PMID: 31068062 Free PMC article. Review.
-
ARF GTPases and their GEFs and GAPs: concepts and challenges.Mol Biol Cell. 2019 May 15;30(11):1249-1271. doi: 10.1091/mbc.E18-12-0820. Mol Biol Cell. 2019. PMID: 31084567 Free PMC article. Review.
-
Rag GTPases Suppress Renal Cystic Disease by Inhibiting TFEB Independently of mTORC1.bioRxiv [Preprint]. 2025 Jul 31:2025.07.24.664930. doi: 10.1101/2025.07.24.664930. bioRxiv. 2025. PMID: 40766400 Free PMC article. Preprint.
-
Probe into the Target and Mechanism of Jianpi Xiaoke Prescription for Treating Type 2 Diabetes Mellitus through miRNA Expression Profiling.Evid Based Complement Alternat Med. 2020 Dec 15;2020:7370350. doi: 10.1155/2020/7370350. eCollection 2020. Evid Based Complement Alternat Med. 2020. PMID: 33456489 Free PMC article.
-
Intrapancreatic fat, pancreatitis, and pancreatic cancer.Cell Mol Life Sci. 2023 Jul 15;80(8):206. doi: 10.1007/s00018-023-04855-z. Cell Mol Life Sci. 2023. PMID: 37452870 Free PMC article. Review.
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases
Research Materials

