MicroRNA expression profiling of Xp11 renal cell carcinoma
- PMID: 28411178
- PMCID: PMC5628161
- DOI: 10.1016/j.humpath.2017.03.011
MicroRNA expression profiling of Xp11 renal cell carcinoma
Abstract
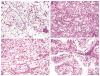
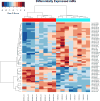
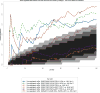
Renal cell carcinomas (RCCs) with Xp11 translocation (Xp11 RCC) constitute a distinctive molecular subtype characterized by chromosomal translocations involving the Xp11.2 locus, resulting in gene fusions between the TFE3 transcription factor with a second gene (usually ASPSCR1, PRCC, NONO, or SFPQ). RCCs with Xp11 translocations comprise up to 1% to 4% of adult cases, frequently displaying papillary architecture with epithelioid clear cells. To better understand the biology of this molecularly distinct tumor subtype, we analyze the microRNA (miRNA) expression profiles of Xp11 RCC compared with normal renal parenchyma using microarray and quantitative reverse-transcription polymerase chain reaction. We further compare Xp11 RCC with other RCC histologic subtypes using publically available data sets, identifying common and distinctive miRNA signatures along with the associated signaling pathways and biological processes. Overall, Xp11 RCC more closely resembles clear cell rather than papillary RCC. Furthermore, among the most differentially expressed miRNAs specific for Xp11 RCC, we identify miR-148a-3p, miR-221-3p, miR-185-5p, miR-196b-5p, and miR-642a-5p to be up-regulated, whereas miR-133b and miR-658 were down-regulated. Finally, Xp11 RCC is most strongly associated with miRNA expression profiles modulating DNA damage responses, cell cycle progression and apoptosis, and the Hedgehog signaling pathway. In summary, we describe here for the first time the miRNA expression profiles of a molecularly distinct type of renal cancer associated with Xp11.2 translocations involving the TFE3 gene. Our results might help understanding the molecular underpinning of Xp11 RCC, assisting in developing targeted treatments for this disease.
Keywords: Analysis of Functional Annotation; Renal cell carcinoma; TFE3 gene fusion; Xp11 translocation; gene regulation; microRNA expression profiling.
Copyright © 2017 Elsevier Inc. All rights reserved.
Conflict of interest statement
Figures


Comment in
-
MicroRNAs, promising biomarkers in the diagnosis of Xp11 translocation RCC.Hum Pathol. 2017 Oct;68:205-206. doi: 10.1016/j.humpath.2017.06.022. Epub 2017 Aug 12. Hum Pathol. 2017. PMID: 28807734 No abstract available.
-
MicroRNAs, promising biomarkers in the diagnosis of Xp11 translocation RCC-reply.Hum Pathol. 2017 Oct;68:206-207. doi: 10.1016/j.humpath.2017.06.021. Epub 2017 Aug 12. Hum Pathol. 2017. PMID: 28811253 No abstract available.
References
-
- Argani P, Antonescu CR, Couturier J, Fournet J-C, Sciot R, Debiec-Rychter M, et al. PRCC-TFE3 renal carcinomas: morphologic, immunohistochemical, ultrastructural, and molecular analysis of an entity associated with the t(X;1)(p11. 2;q21) Am J Surg Pathol. 2002;26:1553–66. - PubMed
-
- Sukov WR, Hodge JC, Lohse CM, Leibovich BC, Thompson RH, Pearce KE, et al. TFE3 rearrangements in adult renal cell carcinoma: clinical and pathologic features with outcome in a large series of consecutively treated patients. Am J Surg Pathol. 2012;36:663–70. doi: 10.1097/PAS.0b013e31824dd972. - DOI - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical

