High-resolution myogenic lineage mapping by single-cell mass cytometry
- PMID: 28414312
- PMCID: PMC5728993
- DOI: 10.1038/ncb3507
High-resolution myogenic lineage mapping by single-cell mass cytometry
Erratum in
-
Publisher Correction: High-resolution myogenic lineage mapping by single-cell mass cytometry.Nat Cell Biol. 2018 Aug;20(8):990. doi: 10.1038/s41556-018-0043-1. Nat Cell Biol. 2018. PMID: 29507406
Abstract
Muscle regeneration is a dynamic process during which cell state and identity change over time. A major roadblock has been a lack of tools to resolve a myogenic progression in vivo. Here we capitalize on a transformative technology, single-cell mass cytometry (CyTOF), to identify in vivo skeletal muscle stem cell and previously unrecognized progenitor populations that precede differentiation. We discovered two cell surface markers, CD9 and CD104, whose combined expression enabled in vivo identification and prospective isolation of stem and progenitor cells. Data analysis using the X-shift algorithm paired with single-cell force-directed layout visualization defined a molecular signature of the activated stem cell state (CD44+/CD98+/MyoD+) and delineated a myogenic trajectory during recovery from acute muscle injury. Our studies uncover the dynamics of skeletal muscle regeneration in vivo and pave the way for the elucidation of the regulatory networks that underlie cell-state transitions in muscle diseases and ageing.
Conflict of interest statement
The authors declare no competing financial interests.
Figures

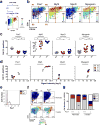




Similar articles
-
Sdf-1 (CXCL12) induces CD9 expression in stem cells engaged in muscle regeneration.Stem Cell Res Ther. 2015 Mar 24;6(1):46. doi: 10.1186/s13287-015-0041-1. Stem Cell Res Ther. 2015. PMID: 25890097 Free PMC article.
-
β4 integrin marks interstitial myogenic progenitor cells in adult murine skeletal muscle.J Histochem Cytochem. 2012 Jan;60(1):31-44. doi: 10.1369/0022155411428991. J Histochem Cytochem. 2012. PMID: 22205679 Free PMC article.
-
Myogenic specification of side population cells in skeletal muscle.J Cell Biol. 2002 Oct 14;159(1):123-34. doi: 10.1083/jcb.200202092. Epub 2002 Oct 14. J Cell Biol. 2002. PMID: 12379804 Free PMC article.
-
Regulation of skeletal muscle stem cell behavior by Pax3 and Pax7.Cold Spring Harb Symp Quant Biol. 2008;73:307-15. doi: 10.1101/sqb.2008.73.006. Epub 2008 Nov 6. Cold Spring Harb Symp Quant Biol. 2008. PMID: 19022756 Review.
-
Epigenetic regulation of skeletal muscle development and differentiation.Subcell Biochem. 2013;61:139-50. doi: 10.1007/978-94-007-4525-4_7. Subcell Biochem. 2013. PMID: 23150250 Review.
Cited by
-
Characterization of Age-Dependent Changes in Skeletal Muscle Repair and Regeneration Using a Mouse Model of Acute Muscle Injury.Methods Mol Biol. 2025;2857:169-180. doi: 10.1007/978-1-0716-4128-6_16. Methods Mol Biol. 2025. PMID: 39348065
-
A Long Journey before Cycling: Regulation of Quiescence Exit in Adult Muscle Satellite Cells.Int J Mol Sci. 2022 Feb 3;23(3):1748. doi: 10.3390/ijms23031748. Int J Mol Sci. 2022. PMID: 35163665 Free PMC article. Review.
-
It takes all kinds: heterogeneity among satellite cells and fibro-adipogenic progenitors during skeletal muscle regeneration.Development. 2021 Nov 1;148(21):dev199861. doi: 10.1242/dev.199861. Epub 2021 Nov 5. Development. 2021. PMID: 34739030 Free PMC article. Review.
-
Activation of JUN in fibroblasts promotes pro-fibrotic programme and modulates protective immunity.Nat Commun. 2020 Jun 3;11(1):2795. doi: 10.1038/s41467-020-16466-4. Nat Commun. 2020. PMID: 32493933 Free PMC article.
-
Non-equivalence of nuclear import among nuclei in multinucleated skeletal muscle cells.J Cell Sci. 2018 Feb 5;131(3):jcs207670. doi: 10.1242/jcs.207670. J Cell Sci. 2018. PMID: 29361530 Free PMC article.
References
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Miscellaneous

