Trypanosoma brucei TbIF1 inhibits the essential F1-ATPase in the infectious form of the parasite
- PMID: 28414727
- PMCID: PMC5407850
- DOI: 10.1371/journal.pntd.0005552
Trypanosoma brucei TbIF1 inhibits the essential F1-ATPase in the infectious form of the parasite
Abstract
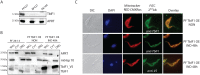
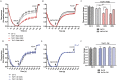
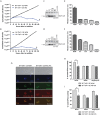
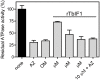
The mitochondrial (mt) FoF1-ATP synthase of the digenetic parasite, Trypanosoma brucei, generates ATP during the insect procyclic form (PF), but becomes a perpetual consumer of ATP in the mammalian bloodstream form (BF), which lacks a canonical respiratory chain. This unconventional dependence on FoF1-ATPase is required to maintain the essential mt membrane potential (Δψm). Normally, ATP hydrolysis by this rotary molecular motor is restricted to when eukaryotic cells experience sporadic hypoxic conditions, during which this compulsory function quickly depletes the cellular ATP pool. To protect against this cellular treason, the highly conserved inhibitory factor 1 (IF1) binds the enzyme in a manner that solely inhibits the hydrolytic activity. Intriguingly, we were able to identify the IF1 homolog in T. brucei (TbIF1), but determined that its expression in the mitochondrion is tightly regulated throughout the life cycle as it is only detected in PF cells. TbIF1 appears to primarily function as an emergency brake in PF cells, where it prevented the restoration of the Δψm by FoF1-ATPase when respiration was chemically inhibited. In vitro, TbIF1 overexpression specifically inhibits the hydrolytic activity but not the synthetic capability of the FoF1-ATP synthase in PF mitochondria. Furthermore, low μM amounts of recombinant TbIF1 achieve the same inhibition of total mt ATPase activity as the FoF1-ATPase specific inhibitors, azide and oligomycin. Therefore, even minimal ectopic expression of TbIF1 in BF cells proved lethal as the indispensable Δψm collapsed due to inhibited FoF1-ATPase. In summary, we provide evidence that T. brucei harbors a natural and potent unidirectional inhibitor of the vital FoF1-ATPase activity that can be exploited for future structure-based drug design.
Conflict of interest statement
The authors have declared that no competing interests exist.
Figures



References
-
- Tielens AG, Van Hellemond JJ (1998) Differences in energy metabolism between trypanosomatidae. Parasitol Today 14: 265–272. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

