Epigenetic regulator CXXC5 recruits DNA demethylase Tet2 to regulate TLR7/9-elicited IFN response in pDCs
- PMID: 28416650
- PMCID: PMC5413332
- DOI: 10.1084/jem.20161149
Epigenetic regulator CXXC5 recruits DNA demethylase Tet2 to regulate TLR7/9-elicited IFN response in pDCs
Abstract
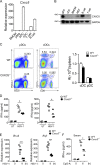
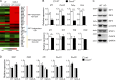
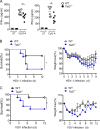
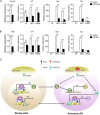
TLR7/9 signals are capable of mounting massive interferon (IFN) response in plasmacytoid dendritic cells (pDCs) immediately after viral infection, yet the involvement of epigenetic regulation in this process has not been documented. Here, we report that zinc finger CXXC family epigenetic regulator CXXC5 is highly expressed in pDCs, where it plays a crucial role in TLR7/9- and virus-induced IFN response. Notably, genetic ablation of CXXC5 resulted in aberrant methylation of the CpG-containing island (CGI) within the Irf7 gene and impaired IRF7 expression in steady-state pDCs. Mechanistically, CXXC5 is responsible for the recruitment of DNA demethylase Tet2 to maintain the hypomethylation of a subset of CGIs, a process coincident with active histone modifications and constitutive transcription of these CGI-containing genes. Consequently, CXXC5-deficient mice had compromised early IFN response and became highly vulnerable to infection by herpes simplex virus and vesicular stomatitis virus. Together, our results identify CXXC5 as a novel epigenetic regulator for pDC-mediated antiviral response.
© 2017 Ma et al.
Figures






Similar articles
-
Spi-B is critical for plasmacytoid dendritic cell function and development.Blood. 2012 Dec 6;120(24):4733-43. doi: 10.1182/blood-2012-06-436527. Epub 2012 Oct 11. Blood. 2012. PMID: 23065153
-
Blimp-1-Mediated Pathway Promotes Type I IFN Production in Plasmacytoid Dendritic Cells by Targeting to Interleukin-1 Receptor-Associated Kinase M.Front Immunol. 2018 Aug 7;9:1828. doi: 10.3389/fimmu.2018.01828. eCollection 2018. Front Immunol. 2018. PMID: 30131810 Free PMC article.
-
Interleukin-1 receptor-associated kinase-1 plays an essential role for Toll-like receptor (TLR)7- and TLR9-mediated interferon-{alpha} induction.J Exp Med. 2005 Mar 21;201(6):915-23. doi: 10.1084/jem.20042372. Epub 2005 Mar 14. J Exp Med. 2005. PMID: 15767370 Free PMC article.
-
CXXC5: A novel regulator and coordinator of TGF-β, BMP and Wnt signaling.J Cell Mol Med. 2019 Feb;23(2):740-749. doi: 10.1111/jcmm.14046. Epub 2018 Nov 27. J Cell Mol Med. 2019. PMID: 30479059 Free PMC article. Review.
-
Plasmacytoid dendritic cells and type I IFN: 50 years of convergent history.Cytokine Growth Factor Rev. 2008 Feb;19(1):3-19. doi: 10.1016/j.cytogfr.2007.10.006. Epub 2008 Jan 11. Cytokine Growth Factor Rev. 2008. PMID: 18248767 Free PMC article. Review.
Cited by
-
Epigenetic Modifiers: Anti-Neoplastic Drugs With Immunomodulating Potential.Front Immunol. 2021 Mar 30;12:652160. doi: 10.3389/fimmu.2021.652160. eCollection 2021. Front Immunol. 2021. PMID: 33859645 Free PMC article. Review.
-
Phenotypical Diversification of Early IFNα-Producing Human Plasmacytoid Dendritic Cells Using Droplet-Based Microfluidics.Front Immunol. 2021 Apr 29;12:672729. doi: 10.3389/fimmu.2021.672729. eCollection 2021. Front Immunol. 2021. PMID: 33995415 Free PMC article.
-
Loss of METTL3 attenuates blastic plasmacytoid dendritic cell neoplasm response to PRMT5 inhibition via IFN signaling.Blood Adv. 2022 Sep 27;6(18):5330-5344. doi: 10.1182/bloodadvances.2021006306. Blood Adv. 2022. PMID: 35482445 Free PMC article.
-
Role of Myeloid Tet Methylcytosine Dioxygenase 2 in Pulmonary and Peritoneal Inflammation Induced by Lipopolysaccharide and Peritonitis Induced by Escherichia coli.Cells. 2021 Dec 28;11(1):82. doi: 10.3390/cells11010082. Cells. 2021. PMID: 35011643 Free PMC article.
-
Tet2 at the interface between cancer and immunity.Commun Biol. 2020 Nov 12;3(1):667. doi: 10.1038/s42003-020-01391-5. Commun Biol. 2020. PMID: 33184433 Free PMC article. Review.
References
-
- Bach P., Kamphuis E., Odermatt B., Sutter G., Buchholz C.J., and Kalinke U.. 2007. Vesicular stomatitis virus glycoprotein displaying retrovirus-like particles induce a type I IFN receptor-dependent switch to neutralizing IgG antibodies. J. Immunol. 178:5839–5847. 10.4049/jimmunol.178.9.5839 - DOI - PubMed
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

