Population Genomics of Legionella longbeachae and Hidden Complexities of Infection Source Attribution
- PMID: 28418314
- PMCID: PMC5403047
- DOI: 10.3201/eid2305.161165
Population Genomics of Legionella longbeachae and Hidden Complexities of Infection Source Attribution
Abstract
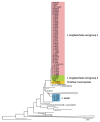
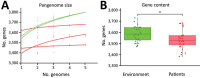
Legionella longbeachae is the primary cause of legionellosis in Australasia and Southeast Asia and an emerging pathogen in Europe and the United States; however, our understanding of the population diversity of L. longbeachae from patient and environmental sources is limited. We analyzed the genomes of 64 L. longbeachae isolates, of which 29 were from a cluster of legionellosis cases linked to commercial growing media in Scotland in 2013 and 35 were non-outbreak-associated isolates from Scotland and other countries. We identified extensive genetic diversity across the L. longbeachae species, associated with intraspecies and interspecies gene flow, and a wide geographic distribution of closely related genotypes. Of note, we observed a highly diverse pool of L. longbeachae genotypes within compost samples that precluded the genetic establishment of an infection source. These data represent a view of the genomic diversity of L. longbeachae that will inform strategies for investigating future outbreaks.
Keywords: Legionella longbeachae; bacteria; environmental genotypes; epidemiology; genomic diversity; legionella; legionellosis; outbreaks; pathogenic genotypes; phylogeny; plasmids; recombination; source attribution.
Figures


Similar articles
-
Legionella longbeachae detected in an industrial cooling tower linked to a legionellosis outbreak, New Zealand, 2015; possible waterborne transmission?Epidemiol Infect. 2017 Aug;145(11):2382-2389. doi: 10.1017/S0950268817001170. Epub 2017 Jun 19. Epidemiol Infect. 2017. PMID: 28625225 Free PMC article.
-
Legionella longbeachae serogroup 1 infections linked to potting compost.J Med Microbiol. 2012 Feb;61(Pt 2):218-222. doi: 10.1099/jmm.0.035857-0. Epub 2011 Sep 22. J Med Microbiol. 2012. PMID: 21940651
-
Combining Environmental Investigation and a Dual-Analytical Strategy to Isolate the Legionella longbeachae Strain Linked to Two Occupational Cases of Legionellosis.Ann Work Expo Health. 2018 Mar 12;62(3):321-327. doi: 10.1093/annweh/wxx109. Ann Work Expo Health. 2018. PMID: 29304227
-
Compost and Legionella longbeachae: an emerging infection?Perspect Public Health. 2015 Nov;135(6):309-15. doi: 10.1177/1757913915611162. Perspect Public Health. 2015. PMID: 26543151 Review.
-
Legionnaires' disease in immunocompromised patients: a case report of Legionella longbeachae pneumonia and review of the literature.J Med Microbiol. 2008 Mar;57(Pt 3):384-387. doi: 10.1099/jmm.0.47556-0. J Med Microbiol. 2008. PMID: 18287305 Review.
Cited by
-
Legionella longbeachae effector protein RavZ inhibits autophagy and regulates phagosome ubiquitination during infection.PLoS One. 2023 Feb 9;18(2):e0281587. doi: 10.1371/journal.pone.0281587. eCollection 2023. PLoS One. 2023. PMID: 36758031 Free PMC article.
-
Epileptic Seizure after Use of Moxifloxacin in Man with Legionella longbeachae Pneumonia.Emerg Infect Dis. 2020 Nov;26(11):2725-2727. doi: 10.3201/eid2611.191815. Emerg Infect Dis. 2020. PMID: 33079050 Free PMC article.
-
Assessing the impact, genomics and evolution of type II secretion across a large, medically important genus: the Legionella type II secretion paradigm.Microb Genom. 2019 Jun;5(6):e000273. doi: 10.1099/mgen.0.000273. Epub 2019 Jun 5. Microb Genom. 2019. PMID: 31166887 Free PMC article.
-
Complete Genome Sequence of Legionella sainthelensi Isolated from a Patient with Legionnaires' Disease.Genome Announc. 2018 Feb 1;6(5):e01588-17. doi: 10.1128/genomeA.01588-17. Genome Announc. 2018. PMID: 29437115 Free PMC article.
-
Legionella spp. All Ears? The Broad Occurrence of Quorum Sensing Elements outside Legionella pneumophila.Genome Biol Evol. 2021 Mar 1;13(3):evab032. doi: 10.1093/gbe/evab032. Genome Biol Evol. 2021. PMID: 33599258 Free PMC article.
References
-
- European Centre for Disease Prevention and Control. Surveillance report. Legionnaires’ disease in Europe, 2010. 2012. [cited 2016 Jul 9]. http://ecdc.europa.eu/en/publications/publications/sur-legionnaires-dise...
-
- Joseph CA, Ricketts KD; European Working Group for Legionella Infections. Legionnaires disease in Europe 2007-2008. Euro Surveill. 2010;15:19493. - PubMed
-
- Li JS, O’Brien ED, Guest C. A review of national legionellosis surveillance in Australia, 1991 to 2000. Commun Dis Intell Q Rep. 2002;26:461–8. - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

