Brain Region and Isoform-Specific Phosphorylation Alters Kalirin SH2 Domain Interaction Sites and Calpain Sensitivity
- PMID: 28418645
- PMCID: PMC5517348
- DOI: 10.1021/acschemneuro.7b00076
Brain Region and Isoform-Specific Phosphorylation Alters Kalirin SH2 Domain Interaction Sites and Calpain Sensitivity
Abstract
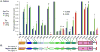
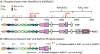
Kalirin7 (Kal7), a postsynaptic Rho GDP/GTP exchange factor (RhoGEF), plays a crucial role in long-term potentiation and in the effects of cocaine on behavior and spine morphology. The KALRN gene has been linked to schizophrenia and other disorders of synaptic function. Mass spectrometry was used to quantify phosphorylation at 26 sites in Kal7 from individual adult rat nucleus accumbens and prefrontal cortex before and after exposure to acute or chronic cocaine. Region- and isoform-specific phosphorylation was observed along with region-specific effects of cocaine on Kal7 phosphorylation. Evaluation of the functional significance of multisite phosphorylation in a complex protein like Kalirin is difficult. With the identification of five tyrosine phosphorylation (pY) sites, a panel of 71 SH2 domains was screened, identifying subsets that interacted with multiple pY sites in Kal7. In addition to this type of reversible interaction, endoproteolytic cleavage by calpain plays an essential role in long-term potentiation. Calpain cleaved Kal7 at two sites, separating the N-terminal domain, which affects spine length, and the PDZ binding motif from the GEF domain. Mutations preventing phosphorylation did not affect calpain sensitivity or GEF activity; phosphomimetic mutations at specific sites altered protein stability, increased calpain sensitivity, and reduced GEF activity.
Keywords: RhoGEF; cocaine; mass spectrometry; nucleus accumbens; prefrontal cortex; tyrosine phosphorylation.
Conflict of interest statement
Figures








Similar articles
-
Kalrn promoter usage and isoform expression respond to chronic cocaine exposure.BMC Neurosci. 2011 Feb 17;12:20. doi: 10.1186/1471-2202-12-20. BMC Neurosci. 2011. PMID: 21329509 Free PMC article.
-
A role for kalirin in the response of rat medium spiny neurons to cocaine.Mol Pharmacol. 2012 Oct;82(4):738-45. doi: 10.1124/mol.112.080044. Epub 2012 Jul 24. Mol Pharmacol. 2012. PMID: 22828798 Free PMC article.
-
Using Kalirin conditional knockout mice to distinguish its role in dopamine receptor mediated behaviors.BMC Neurosci. 2017 May 23;18(1):45. doi: 10.1186/s12868-017-0363-2. BMC Neurosci. 2017. PMID: 28535798 Free PMC article.
-
Kalirin-7 is a key player in the formation of excitatory synapses in hippocampal neurons.ScientificWorldJournal. 2010 Aug 17;10:1655-66. doi: 10.1100/tsw.2010.148. ScientificWorldJournal. 2010. PMID: 20730383 Free PMC article. Review.
-
Kalirin, a key player in synapse formation, is implicated in human diseases.Neural Plast. 2012;2012:728161. doi: 10.1155/2012/728161. Epub 2012 Apr 3. Neural Plast. 2012. PMID: 22548195 Free PMC article. Review.
Cited by
-
Kalirin/Trio Rho GDP/GTP exchange factors regulate proinsulin and insulin secretion.J Mol Endocrinol. 2019 Jan;62(1):47-65. doi: 10.1530/JME-18-0048. J Mol Endocrinol. 2019. PMID: 30407917 Free PMC article.
-
Role of Kalirin and mouse strain in retention of spatial memory training in an Alzheimer's disease model mouse line.Neurobiol Aging. 2020 Nov;95:69-80. doi: 10.1016/j.neurobiolaging.2020.07.006. Epub 2020 Jul 14. Neurobiol Aging. 2020. PMID: 32768866 Free PMC article.
-
Getting Started with the IDG KMC Datasets and Tools.Curr Protoc. 2022 Jan;2(1):e355. doi: 10.1002/cpz1.355. Curr Protoc. 2022. PMID: 35085427 Free PMC article.
-
CELA2A mutations predispose to early-onset atherosclerosis and metabolic syndrome and affect plasma insulin and platelet activation.Nat Genet. 2019 Aug;51(8):1233-1243. doi: 10.1038/s41588-019-0470-3. Epub 2019 Jul 29. Nat Genet. 2019. PMID: 31358993 Free PMC article.
-
Regulated processing and secretion of a peptide precursor in cilia.Proc Natl Acad Sci U S A. 2022 Aug 2;119(31):e2206098119. doi: 10.1073/pnas.2206098119. Epub 2022 Jul 25. Proc Natl Acad Sci U S A. 2022. PMID: 35878031 Free PMC article.
References
-
- Hayashi-Takagi A, Takaki M, Graziane N, Seshadri S, Murdoch H, Dunlop AJ, Makino Y, Seshadri AJ, Ishizuka K, Srivastava DP, Xie Z, Baraban JM, Houslay MD, Tomoda T, Brandon NJ, Kamiya A, Yan Z, Penzes P, Sawa A. Disrupted-in-Schizophrenia 1 (DISC1) regulates spines of the glutamate synapse via Rac1. Nature Neurosci. 2010;13:327–332. - PMC - PubMed
-
- Hill JJ, Hashimoto T, Lewis DA. Molecular mechanisms contributing to dendritic spine alterations in the prefrontal cortex of subjects with schizophrenia. Mol Psychiatry. 2006;11:557–566. - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

