Therapeutic inhibition of USP7-PTEN network in chronic lymphocytic leukemia: a strategy to overcome TP53 mutated/deleted clones
- PMID: 28418900
- PMCID: PMC5482594
- DOI: 10.18632/oncotarget.16348
Therapeutic inhibition of USP7-PTEN network in chronic lymphocytic leukemia: a strategy to overcome TP53 mutated/deleted clones
Abstract
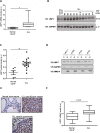
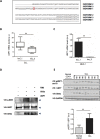
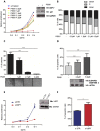
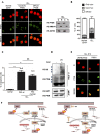
Chronic Lymphocytic Leukemia (CLL) is a lymphoproliferative disorder with either indolent or aggressive clinical course. Current treatment regiments have significantly improved the overall outcomes even if higher risk subgroups - those harboring TP53 mutations or deletions of the short arm of chromosome 17 (del17p) - remain highly challenging. In the present work, we identified USP7, a known de-ubiquitinase with multiple roles in cellular homeostasis, as a potential therapeutic target in CLL. We demonstrated that in primary CLL samples and in CLL cell lines USP7 is: i) over-expressed through a mechanism involving miR-338-3p and miR-181b deregulation; ii) functionally activated by Casein Kinase 2 (CK2), an upstream interactor known to be deregulated in CLL; iii) effectively targeted by the USP7 inhibitor P5091. Treatment of primary CLL samples and cell lines with P5091 induces cell growth arrest and apoptosis, through the restoration of PTEN nuclear pool, both in TP53-wild type and -null environment. Importantly, PTEN acts as the main tumor suppressive mediator along the USP7-PTEN axis in a p53 dispensable manner. In conclusion, we propose USP7 as a new druggable target in CLL.
Keywords: PTEN; USP7; chronic lymphocytic leukemia; miR181; miR338.
Conflict of interest statement
The authors declare that they have no conflict of interest.
Figures

Comment in
-
A laboratory test of evolutionary aging theories.Aging (Albany NY). 2017 Mar 21;9(3):600-601. doi: 10.18632/aging.101215. Aging (Albany NY). 2017. PMID: 28325887 Free PMC article. No abstract available.
-
Tumor suppressors revival in CLL.Aging (Albany NY). 2017 Jun 26;9(6):1473-1474. doi: 10.18632/aging.101258. Aging (Albany NY). 2017. PMID: 28657542 Free PMC article. No abstract available.
Similar articles
-
USP7 inhibition alters homologous recombination repair and targets CLL cells independently of ATM/p53 functional status.Blood. 2017 Jul 13;130(2):156-166. doi: 10.1182/blood-2016-12-758219. Epub 2017 May 11. Blood. 2017. PMID: 28495793
-
The USP7 Inhibitor P5091 Induces Cell Death in Ovarian Cancers with Different P53 Status.Cell Physiol Biochem. 2017;43(5):1755-1766. doi: 10.1159/000484062. Epub 2017 Oct 19. Cell Physiol Biochem. 2017. PMID: 29049989
-
miR-26a and miR-214 down-regulate expression of the PTEN gene in chronic lymphocytic leukemia, but not PTEN mutation or promoter methylation.Oncotarget. 2015 Jan 20;6(2):1276-85. doi: 10.18632/oncotarget.2626. Oncotarget. 2015. PMID: 25361012 Free PMC article.
-
Prognostic and therapeutic stratification in CLL: focus on 17p deletion and p53 mutation.Ann Hematol. 2018 Dec;97(12):2269-2278. doi: 10.1007/s00277-018-3503-6. Epub 2018 Oct 12. Ann Hematol. 2018. PMID: 30315344 Review.
-
The role of TP53 network in the pathogenesis of chronic lymphocytic leukemia.Int J Clin Exp Pathol. 2013 Jun 15;6(7):1223-9. Print 2013. Int J Clin Exp Pathol. 2013. PMID: 23826404 Free PMC article. Review.
Cited by
-
[Expression of ubiquitin-specific protease 7 in lung tissue of preterm rats after hyperoxia exposure].Zhongguo Dang Dai Er Ke Za Zhi. 2020 Dec;22(12):1331-1337. doi: 10.7499/j.issn.1008-8830.2007147. Zhongguo Dang Dai Er Ke Za Zhi. 2020. PMID: 33328006 Free PMC article. Chinese.
-
Targeting casein kinase 2 and ubiquitin-specific protease 7 to modulate RUNX2-mediated osteogenesis in chronic kidney disease.Mol Med. 2025 May 30;31(1):214. doi: 10.1186/s10020-025-01222-5. Mol Med. 2025. PMID: 40447997 Free PMC article.
-
USP7: Novel Drug Target in Cancer Therapy.Front Pharmacol. 2019 Apr 30;10:427. doi: 10.3389/fphar.2019.00427. eCollection 2019. Front Pharmacol. 2019. PMID: 31114498 Free PMC article.
-
Deubiquitinases in hematological malignancies.Biomark Res. 2021 Aug 28;9(1):66. doi: 10.1186/s40364-021-00320-w. Biomark Res. 2021. PMID: 34454635 Free PMC article. Review.
-
Tumor suppressors revival in CLL.Aging (Albany NY). 2017 Jun 26;9(6):1473-1474. doi: 10.18632/aging.101258. Aging (Albany NY). 2017. PMID: 28657542 Free PMC article. No abstract available.
References
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials
Miscellaneous

