Effect of genomic distance on coexpression of coregulated genes in E. coli
- PMID: 28419102
- PMCID: PMC5395161
- DOI: 10.1371/journal.pone.0174887
Effect of genomic distance on coexpression of coregulated genes in E. coli
Abstract
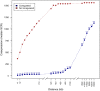
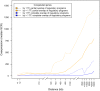
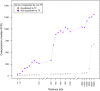
In prokaryotes, genomic distance is a feature that in addition to coregulation affects coexpression. Several observations, such as genomic clustering of highly coexpressed small regulons, support the idea that coexpression behavior of coregulated genes is affected by the distance between the coregulated genes. However, the specific contribution of distance in addition to coregulation in determining the degree of coexpression has not yet been studied systematically. In this work, we exploit the rich information in RegulonDB to study how the genomic distance between coregulated genes affects their degree of coexpression, measured by pairwise similarity of expression profiles obtained under a large number of conditions. We observed that, in general, coregulated genes display higher degrees of coexpression as they are more closely located on the genome. This contribution of genomic distance in determining the degree of coexpression was relatively small compared to the degree of coexpression that was determined by the tightness of the coregulation (degree of overlap of regulatory programs) but was shown to be evolutionary constrained. In addition, the distance effect was sufficient to guarantee coexpression of coregulated genes that are located at very short distances, irrespective of their tightness of coregulation. This is partly but definitely not always because the close distance is also the cause of the coregulation. In cases where it is not, we hypothesize that the effect of the distance on coexpression could be caused by the fact that coregulated genes closely located to each other are also relatively more equidistantly located from their common TF and therefore subject to more similar levels of TF molecules. The absolute genomic distance of the coregulated genes to their common TF-coding gene tends to be less important in determining the degree of coexpression. Our results pinpoint the importance of taking into account the combined effect of distance and coregulation when studying prokaryotic coexpression and transcriptional regulation.
Conflict of interest statement
Figures

Similar articles
-
Interpreting patterns of gene expression: signatures of coregulation, the data processing inequality, and triplet motifs.PLoS One. 2012;7(2):e31969. doi: 10.1371/journal.pone.0031969. Epub 2012 Feb 29. PLoS One. 2012. PMID: 22393375 Free PMC article.
-
Functionally Related Genes Cluster into Genomic Regions That Coordinate Transcription at a Distance in Saccharomyces cerevisiae.mSphere. 2019 Mar 13;4(2):e00063-19. doi: 10.1128/mSphere.00063-19. mSphere. 2019. PMID: 30867326 Free PMC article.
-
Minimal effect of gene clustering on expression in Escherichia coli.Genetics. 2013 Feb;193(2):453-65. doi: 10.1534/genetics.112.147199. Epub 2012 Dec 5. Genetics. 2013. PMID: 23222655 Free PMC article.
-
Prokaryotic genome regulation: multifactor promoters, multitarget regulators and hierarchic networks.FEMS Microbiol Rev. 2010 Sep;34(5):628-45. doi: 10.1111/j.1574-6976.2010.00227.x. Epub 2010 Apr 14. FEMS Microbiol Rev. 2010. PMID: 20491932 Review.
-
Leucine-responsive regulatory protein: a global regulator of gene expression in E. coli.Annu Rev Microbiol. 1995;49:747-75. doi: 10.1146/annurev.mi.49.100195.003531. Annu Rev Microbiol. 1995. PMID: 8561478 Review.
Cited by
-
Redefining fundamental concepts of transcription initiation in bacteria.Nat Rev Genet. 2020 Nov;21(11):699-714. doi: 10.1038/s41576-020-0254-8. Epub 2020 Jul 14. Nat Rev Genet. 2020. PMID: 32665585 Free PMC article. Review.
-
Using RegulonDB, the Escherichia coli K-12 Gene Regulatory Transcriptional Network Database.Curr Protoc Bioinformatics. 2018 Mar;61(1):1.32.1-1.32.30. doi: 10.1002/cpbi.43. Curr Protoc Bioinformatics. 2018. PMID: 30040192 Free PMC article.
-
DNA supercoiling-mediated collective behavior of co-transcribing RNA polymerases.Nucleic Acids Res. 2022 Feb 22;50(3):1269-1279. doi: 10.1093/nar/gkab1252. Nucleic Acids Res. 2022. PMID: 34951454 Free PMC article.
-
BINDER: computationally inferring a gene regulatory network for Mycobacterium abscessus.BMC Bioinformatics. 2019 Sep 10;20(1):466. doi: 10.1186/s12859-019-3042-8. BMC Bioinformatics. 2019. PMID: 31500560 Free PMC article.
-
Stable Housekeeping Genes in Bone Marrow, Adipose Tissue, and Amniotic Membrane-Derived Mesenchymal Stromal Cells for Orthopedic Regenerative Medicine Approaches.Int J Mol Sci. 2024 Jan 25;25(3):1461. doi: 10.3390/ijms25031461. Int J Mol Sci. 2024. PMID: 38338737 Free PMC article.
References
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

