Effects of Active Site Mutations on Specificity of Nucleobase Binding in Human DNA Polymerase η
- PMID: 28423907
- PMCID: PMC5402696
- DOI: 10.1021/acs.jpcb.6b09973
Effects of Active Site Mutations on Specificity of Nucleobase Binding in Human DNA Polymerase η
Abstract
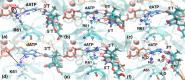
Human DNA polymerase η (Pol η) plays a vital role in protection against skin cancer caused by damage from ultraviolet light. This enzyme rescues stalled replication forks at cyclobutane thymine-thymine dimers (TTDs) by inserting nucleotides opposite these DNA lesions. Residue R61 is conserved in the Pol η enzymes across species, but the corresponding residue, as well as its neighbor S62, is different in other Y-family polymerases, Pol ι and Pol κ. Herein, R61 and S62 are mutated to their Pol ι and Pol κ counterparts. Relative binding free energies of dATP to mutant Pol η•DNA complexes with and without a TTD were calculated using thermodynamic integration. The binding free energies of dATP to the Pol η•DNA complex with and without a TTD are more similar for all of these mutants than for wild-type Pol η, suggesting that these mutations decrease the ability of this enzyme to distinguish between a TTD lesion and undamaged DNA. Molecular dynamics simulations of the mutant systems provide insights into the molecular level basis for the changes in relative binding free energies. The simulations identified differences in hydrogen-bonding, cation-π, and π-π interactions of the side chains with the dATP and the TTD or thymine-thymine (TT) motif. The simulations also revealed that R61 and Q38 act as a clamp to position the dATP and the TTD or TT and that the mutations impact the balance among the interactions related to this clamp. Overall, these calculations suggest that R61 and S62 play key roles in the specificity and effectiveness of Pol η for bypassing TTD lesions during DNA replication. Understanding the basis for this specificity is important for designing drugs aimed at cancer treatment.
Conflict of interest statement
The authors declare no competing financial interest.
Figures
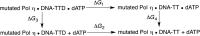




References
-
- Cruet-Hennequart S.; Gallagher K.; Sokòl A.; Villalan S.; Prendergast Á.; Carty M.. DNA polymerase η, a key protein in translesion synthesis in human cells. In Genome Stability and Human Diseases; Nasheuer H.-P., Ed.; Springer: Dordrecht, The Netherlands, 2010; Vol. 50, pp 189–209. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials
Miscellaneous

