MuPeXI: prediction of neo-epitopes from tumor sequencing data
- PMID: 28429069
- PMCID: PMC11028452
- DOI: 10.1007/s00262-017-2001-3
MuPeXI: prediction of neo-epitopes from tumor sequencing data
Abstract
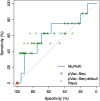
Personalization of immunotherapies such as cancer vaccines and adoptive T cell therapy depends on identification of patient-specific neo-epitopes that can be specifically targeted. MuPeXI, the mutant peptide extractor and informer, is a program to identify tumor-specific peptides and assess their potential to be neo-epitopes. The program input is a file with somatic mutation calls, a list of HLA types, and optionally a gene expression profile. The output is a table with all tumor-specific peptides derived from nucleotide substitutions, insertions, and deletions, along with comprehensive annotation, including HLA binding and similarity to normal peptides. The peptides are sorted according to a priority score which is intended to roughly predict immunogenicity. We applied MuPeXI to three tumors for which predicted MHC-binding peptides had been screened for T cell reactivity, and found that MuPeXI was able to prioritize immunogenic peptides with an area under the curve of 0.63. Compared to other available tools, MuPeXI provides more information and is easier to use. MuPeXI is available as stand-alone software and as a web server at http://www.cbs.dtu.dk/services/MuPeXI .
Keywords: Immunotherapy; Mutation; Neo-antigens; Neo-epitopes; Prediction; Sequencing.
Conflict of interest statement
The authors declare that they have no conflict of interest.
Figures

Similar articles
-
A Computational Pipeline for Predicting Cancer Neoepitopes.Methods Mol Biol. 2023;2552:475-488. doi: 10.1007/978-1-0716-2609-2_27. Methods Mol Biol. 2023. PMID: 36346610
-
Level of neo-epitope predecessor and mutation type determine T cell activation of MHC binding peptides.J Immunother Cancer. 2019 May 22;7(1):135. doi: 10.1186/s40425-019-0595-z. J Immunother Cancer. 2019. PMID: 31118084 Free PMC article.
-
Prediction of neo-epitope immunogenicity reveals TCR recognition determinants and provides insight into immunoediting.Cell Rep Med. 2021 Feb 6;2(2):100194. doi: 10.1016/j.xcrm.2021.100194. eCollection 2021 Feb 16. Cell Rep Med. 2021. PMID: 33665637 Free PMC article.
-
Recent Advances in Lung Cancer Immunotherapy: Input of T-Cell Epitopes Associated With Impaired Peptide Processing.Front Immunol. 2019 Jul 3;10:1505. doi: 10.3389/fimmu.2019.01505. eCollection 2019. Front Immunol. 2019. PMID: 31333652 Free PMC article. Review.
-
Targeting Neoepitopes to Treat Solid Malignancies: Immunosurgery.Front Immunol. 2021 Jul 15;12:592031. doi: 10.3389/fimmu.2021.592031. eCollection 2021. Front Immunol. 2021. PMID: 34335558 Free PMC article. Review.
Cited by
-
Evaluation of T-Cell Responses Against Shared Melanoma Associated Antigens and Predicted Neoantigens in Cutaneous Melanoma Patients Treated With the CSF-470 Allogeneic Cell Vaccine Plus BCG and GM-CSF.Front Immunol. 2020 Jun 5;11:1147. doi: 10.3389/fimmu.2020.01147. eCollection 2020. Front Immunol. 2020. PMID: 32582212 Free PMC article. Clinical Trial.
-
The Landscape of Tumor Fusion Neoantigens: A Pan-Cancer Analysis.iScience. 2019 Nov 22;21:249-260. doi: 10.1016/j.isci.2019.10.028. Epub 2019 Oct 18. iScience. 2019. PMID: 31677477 Free PMC article.
-
pVACtools: A Computational Toolkit to Identify and Visualize Cancer Neoantigens.Cancer Immunol Res. 2020 Mar;8(3):409-420. doi: 10.1158/2326-6066.CIR-19-0401. Epub 2020 Jan 6. Cancer Immunol Res. 2020. PMID: 31907209 Free PMC article.
-
Genomic predictors of response to PD-1 inhibition in children with germline DNA replication repair deficiency.Nat Med. 2022 Jan;28(1):125-135. doi: 10.1038/s41591-021-01581-6. Epub 2022 Jan 6. Nat Med. 2022. PMID: 34992263 Free PMC article.
-
SNPnexus: assessing the functional relevance of genetic variation to facilitate the promise of precision medicine.Nucleic Acids Res. 2018 Jul 2;46(W1):W109-W113. doi: 10.1093/nar/gky399. Nucleic Acids Res. 2018. PMID: 29757393 Free PMC article.
References
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials

