Arf GAPs and molecular motors
- PMID: 28430047
- PMCID: PMC6548297
- DOI: 10.1080/21541248.2017.1308850
Arf GAPs and molecular motors
Abstract
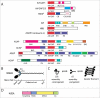
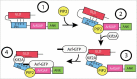
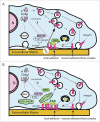
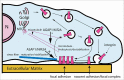
Arf GTPase-activating proteins (Arf GAPs) were first identified as regulators of the small GTP-binding proteins ADP-ribosylation factors (Arfs). The Arf GAPs are a large family of proteins in metazoans, outnumbering the Arfs that they regulate. The members of the Arf GAP family have complex domain structures and some have been implicated in particular cellular functions, such as cell migration, or with particular pathologies, such as tumor invasion and metastasis. The specific effects of Arfs sometimes depend on the Arf GAP involved in their regulation. These observations have led to speculation that the Arf GAPs themselves may affect cellular activities in capacities beyond the regulation of Arfs. Recently, 2 Arf GAPs, ASAP1 and AGAP1, have been found to bind directly to and influence the activity of myosins and kinesins, motor proteins associated with filamentous actin and microtubules, respectively. The Arf GAP-motor protein interaction is critical for cellular behaviors involving the actin cytoskeleton and microtubules, such as cell migration and other cell movements. Arfs, then, may function with molecular motors through Arf GAPs to regulate microtubule and actin remodeling.
Keywords: ADP-ribosylation factor GTPase-activating protein; ADP-ribosylation factors; AGAP1; ASAP1; Arf; Arf GAP; Kif2A; NM2A; kinesin-13; nonmuscle myosin 2A.
Figures


Similar articles
-
Arf GAPs as Regulators of the Actin Cytoskeleton-An Update.Int J Mol Sci. 2019 Jan 21;20(2):442. doi: 10.3390/ijms20020442. Int J Mol Sci. 2019. PMID: 30669557 Free PMC article. Review.
-
Direct Functional Interaction of the Kinesin-13 Family Member Kinesin-like Protein 2A (Kif2A) and Arf GAP with GTP-binding Protein-like, Ankyrin Repeats and PH Domains1 (AGAP1).J Biol Chem. 2016 Oct 7;291(41):21350-21362. doi: 10.1074/jbc.M116.732479. Epub 2016 Aug 16. J Biol Chem. 2016. PMID: 27531749 Free PMC article.
-
Differences between AGAP1, ASAP1 and Arf GAP1 in substrate recognition: interaction with the N-terminus of Arf1.Cell Signal. 2004 Sep;16(9):1033-44. doi: 10.1016/j.cellsig.2004.02.008. Cell Signal. 2004. PMID: 15212764
-
Assays and properties of the Arf GAPs AGAP1, ASAP1, and Arf GAP1.Methods Enzymol. 2005;404:147-63. doi: 10.1016/S0076-6879(05)04015-2. Methods Enzymol. 2005. PMID: 16413266
-
Arf GAPs as regulators of the actin cytoskeleton.Biol Cell. 2007 Oct;99(10):583-600. doi: 10.1042/bc20070034. Biol Cell. 2007. PMID: 17868031 Review.
Cited by
-
ARF GTPases and their GEFs and GAPs: concepts and challenges.Mol Biol Cell. 2019 May 15;30(11):1249-1271. doi: 10.1091/mbc.E18-12-0820. Mol Biol Cell. 2019. PMID: 31084567 Free PMC article. Review.
-
Parkinson's disease-associated alterations in DNA methylation and hydroxymethylation in human brain.bioRxiv [Preprint]. 2025 Feb 2:2024.05.21.595193. doi: 10.1101/2024.05.21.595193. bioRxiv. 2025. PMID: 39975085 Free PMC article. Preprint.
-
Arf GAPs as Regulators of the Actin Cytoskeleton-An Update.Int J Mol Sci. 2019 Jan 21;20(2):442. doi: 10.3390/ijms20020442. Int J Mol Sci. 2019. PMID: 30669557 Free PMC article. Review.
-
Stratifying Cumulus Cell Samples Based on Molecular Profiling to Help Resolve Biomarker Discrepancies and to Predict Oocyte Developmental Competence.Int J Mol Sci. 2021 Jun 15;22(12):6377. doi: 10.3390/ijms22126377. Int J Mol Sci. 2021. PMID: 34203623 Free PMC article.
-
Role of oxidative stress in Retinitis pigmentosa: new involved pathways by an RNA-Seq analysis.Cell Cycle. 2019 Jan;18(1):84-104. doi: 10.1080/15384101.2018.1558873. Epub 2018 Dec 28. Cell Cycle. 2019. Retraction in: Cell Cycle. 2020 Jan;19(2):256. doi: 10.1080/15384101.2019.1702311. PMID: 30569795 Free PMC article. Retracted.
References
-
- Kahn RA, Cherfils J, Elias M, Lovering RC, Munro S, Schurmann A. Nomenclature for the human Arf family of GTP-binding proteins: ARF, ARL, and SAR proteins. J Cell Biol 2006; 172:645–50; PMID:16505163; http://dx.doi.org/10.1083/jcb.200512057 - DOI - PMC - PubMed
-
- Gillingham AK, Munro S. The small G proteins of the arf family and their regulators. Ann Rev Cell Dev Biol 2007; 23:579–611; http://dx.doi.org/10.1146/annurev.cellbio.23.090506.123209 - DOI - PubMed
-
- D'Souza-Schorey C, Chavrier P. ARF proteins: roles in membrane traffic and beyond. Nat Rev Mol Cell Biol 2006; 7:347–58; PMID:16633337; http://dx.doi.org/10.1038/nrm1910 - DOI - PubMed
-
- Donaldson JG, Jackson CL. ARF family G proteins and their regulators: roles in membrane transport, development and disease. Nat Rev Mol Cell Biol 2011; 12:362–75; PMID:21587297; http://dx.doi.org/10.1038/nrm3117 - DOI - PMC - PubMed
-
- Kahn RA, Bruford E, Inoue H, Logsdon JM, Nie ZZ, Premont RT, Randazzo PA, Satake M, Theibert AB, Zapp ML, et al. . Consensus nomenclature for the human ArfGAP domain-containing proteins. J Cell Biol 2008; 182:1039–44; PMID:18809720; http://dx.doi.org/10.1083/jcb.200806041 - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials
Miscellaneous
