A bacterial ABC transporter enables import of mammalian host glycosaminoglycans
- PMID: 28432302
- PMCID: PMC5430744
- DOI: 10.1038/s41598-017-00917-y
A bacterial ABC transporter enables import of mammalian host glycosaminoglycans
Abstract
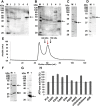
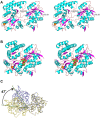
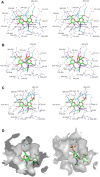
Glycosaminoglycans (GAGs), such as hyaluronan, chondroitin sulfate, and heparin, constitute mammalian extracellular matrices. The uronate and amino sugar residues in hyaluronan and chondroitin sulfate are linked by 1,3-glycoside bond, while heparin contains 1,4-glycoside bond. Some bacteria target GAGs as means of establishing colonization and/or infection, and bacterial degradation mechanisms of GAGs have been well characterized. However, little is known about the bacterial import of GAGs. Here, we show a GAG import system, comprised of a solute-binding protein (Smon0123)-dependent ATP-binding cassette (ABC) transporter, in the pathogenic Streptobacillus moniliformis. A genetic cluster responsible for depolymerization, degradation, and metabolism of GAGs as well as the ABC transporter system was found in the S. moniliformis genome. This bacterium degraded hyaluronan and chondroitin sulfate with an expression of the genetic cluster, while heparin repressed the bacterial growth. The purified recombinant Smon0123 exhibited an affinity with disaccharides generated from hyaluronan and chondroitin sulfate. X-ray crystallography indicated binding mode of Smon0123 to GAG disaccharides. The purified recombinant ABC transporter as a tetramer (Smon0121-Smon0122/Smon0120-Smon0120) reconstructed in liposomes enhanced its ATPase activity in the presence of Smon0123 and GAG disaccharides. This is the first report that has molecularly depicted a bacterial import system of both sulfated and non-sulfated GAGs.
Conflict of interest statement
The authors declare that they have no competing interests.
Figures




Similar articles
-
Streptococcal phosphotransferase system imports unsaturated hyaluronan disaccharide derived from host extracellular matrices.PLoS One. 2019 Nov 7;14(11):e0224753. doi: 10.1371/journal.pone.0224753. eCollection 2019. PLoS One. 2019. PMID: 31697725 Free PMC article.
-
Substrate recognition by bacterial solute-binding protein is responsible for import of extracellular hyaluronan and chondroitin sulfate from the animal host.Biosci Biotechnol Biochem. 2019 Oct;83(10):1946-1954. doi: 10.1080/09168451.2019.1630250. Epub 2019 Jun 16. Biosci Biotechnol Biochem. 2019. PMID: 31204616
-
Alternative substrate-bound conformation of bacterial solute-binding protein involved in the import of mammalian host glycosaminoglycans.Sci Rep. 2017 Dec 5;7(1):17005. doi: 10.1038/s41598-017-16801-8. Sci Rep. 2017. PMID: 29208901 Free PMC article.
-
Implementation of infrared and Raman modalities for glycosaminoglycan characterization in complex systems.Glycoconj J. 2017 Jun;34(3):309-323. doi: 10.1007/s10719-016-9743-6. Epub 2016 Dec 7. Glycoconj J. 2017. PMID: 27928742 Free PMC article. Review.
-
Glycosaminoglycans: Carriers and Targets for Tailored Anti-Cancer Therapy.Biomolecules. 2021 Mar 8;11(3):395. doi: 10.3390/biom11030395. Biomolecules. 2021. PMID: 33800172 Free PMC article. Review.
Cited by
-
Probiotics in human gut microbiota can degrade host glycosaminoglycans.Sci Rep. 2018 Jul 13;8(1):10674. doi: 10.1038/s41598-018-28886-w. Sci Rep. 2018. PMID: 30006634 Free PMC article.
-
The Fish Pathogen Aliivibrio salmonicida LFI1238 Can Degrade and Metabolize Chitin despite Gene Disruption in the Chitinolytic Pathway.Appl Environ Microbiol. 2021 Sep 10;87(19):e0052921. doi: 10.1128/AEM.00529-21. Epub 2021 Sep 10. Appl Environ Microbiol. 2021. PMID: 34319813 Free PMC article.
-
Hungatella hathewayi, an Efficient Glycosaminoglycan-Degrading Firmicutes from Human Gut and Its Chondroitin ABC Exolyase with High Activity and Broad Substrate Specificity.Appl Environ Microbiol. 2022 Nov 22;88(22):e0154622. doi: 10.1128/aem.01546-22. Epub 2022 Nov 7. Appl Environ Microbiol. 2022. PMID: 36342199 Free PMC article.
-
Predictive Metagenomic Profiling, Urine Metabolomics, and Human Marker Gene Expression as an Integrated Approach to Study Alopecia Areata.Front Cell Infect Microbiol. 2020 Apr 29;10:146. doi: 10.3389/fcimb.2020.00146. eCollection 2020. Front Cell Infect Microbiol. 2020. PMID: 32411613 Free PMC article.
-
Streptococcal phosphotransferase system imports unsaturated hyaluronan disaccharide derived from host extracellular matrices.PLoS One. 2019 Nov 7;14(11):e0224753. doi: 10.1371/journal.pone.0224753. eCollection 2019. PLoS One. 2019. PMID: 31697725 Free PMC article.
References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

