Early vertebrate origin and diversification of small transmembrane regulators of cellular ion transport
- PMID: 28436536
- PMCID: PMC5509871
- DOI: 10.1113/JP274254
Early vertebrate origin and diversification of small transmembrane regulators of cellular ion transport
Abstract
Key points: Small transmembrane proteins such as FXYDs, which interact with Na+ ,K+ -ATPase, and the micropeptides that interact with sarco/endoplasmic reticulum Ca2+ -ATPase play fundamental roles in regulation of ion transport in vertebrates. Uncertain evolutionary origins and phylogenetic relationships among these regulators of ion transport have led to inconsistencies in their classification across vertebrate species, thus hampering comparative studies of their functions. We discovered the first FXYD homologue in sea lamprey, a basal jawless vertebrate, which suggests small transmembrane regulators of ion transport emerged early in the vertebrate lineage. We also identified 13 gene subfamilies of FXYDs and propose a revised, phylogeny-based FXYD classification that is consistent across vertebrate species. These findings provide an improved framework for investigating physiological and pathophysiological functions of small transmembrane regulators of ion transport.
Abstract: Small transmembrane proteins are important for regulation of cellular ion transport. The most prominent among these are members of the FXYD family (FXYD1-12), which regulate Na+ ,K+ -ATPase, and phospholamban, sarcolipin, myoregulin and DWORF, which regulate the sarco/endoplasmic reticulum Ca2+ -ATPase (SERCA). FXYDs and regulators of SERCA are present in fishes, as well as terrestrial vertebrates; however, their evolutionary origins and phylogenetic relationships are obscure, thus hampering comparative physiological studies. Here we discovered that sea lamprey (Petromyzon marinus), a representative of extant jawless vertebrates (Cyclostomata), expresses an FXYD homologue, which strongly suggests that FXYDs predate the emergence of fishes and other jawed vertebrates (Gnathostomata). Using a combination of sequence-based phylogenetic analysis and conservation of local chromosome context, we determined that FXYDs markedly diversified in the lineages leading to cartilaginous fishes (Chondrichthyes) and bony vertebrates (Euteleostomi). Diversification of SERCA regulators was much less extensive, indicating they operate under different evolutionary constraints. Finally, we found that FXYDs in extant vertebrates can be classified into 13 gene subfamilies, which do not always correspond to the established FXYD classification. We therefore propose a revised classification that is based on evolutionary history of FXYDs and that is consistent across vertebrate species. Collectively, our findings provide an improved framework for investigating the function of ion transport in health and disease.
Keywords: FXYD proteins; Na+−K+−ATPase; SERCA; micropeptides; phospholemman; protein phosphorylation; vertebrate evolution.
© 2017 The Authors. The Journal of Physiology © 2017 The Physiological Society.
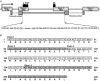
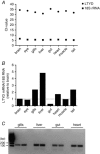
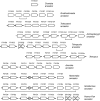
Figures






Similar articles
-
Hormonal regulation of Na+-K+-ATPase from the evolutionary perspective.Curr Top Membr. 2019;83:315-351. doi: 10.1016/bs.ctm.2019.01.009. Epub 2019 Mar 5. Curr Top Membr. 2019. PMID: 31196608 Review.
-
Genome-wide analysis of the ATP-binding cassette (ABC) transporter gene family in sea lamprey and Japanese lamprey.BMC Genomics. 2015 Jun 6;16(1):436. doi: 10.1186/s12864-015-1677-z. BMC Genomics. 2015. PMID: 26047617 Free PMC article.
-
The globin gene repertoire of lampreys: convergent evolution of hemoglobin and myoglobin in jawed and jawless vertebrates.Mol Biol Evol. 2014 Oct;31(10):2708-21. doi: 10.1093/molbev/msu216. Epub 2014 Jul 23. Mol Biol Evol. 2014. PMID: 25061084 Free PMC article.
-
Evolution of hematopoiesis: Three members of the PU.1 transcription factor family in a cartilaginous fish, Raja eglanteria.Proc Natl Acad Sci U S A. 2001 Jan 16;98(2):553-8. doi: 10.1073/pnas.98.2.553. Epub 2001 Jan 9. Proc Natl Acad Sci U S A. 2001. PMID: 11149949 Free PMC article.
-
Functional genetic analysis in a jawless vertebrate, the sea lamprey: insights into the developmental evolution of early vertebrates.J Exp Biol. 2020 Feb 7;223(Pt Suppl 1):jeb206433. doi: 10.1242/jeb.206433. J Exp Biol. 2020. PMID: 32034037 Review.
Cited by
-
FXYD proteins and sodium pump regulatory mechanisms.J Gen Physiol. 2021 Apr 5;153(4):e202012633. doi: 10.1085/jgp.202012633. J Gen Physiol. 2021. PMID: 33688925 Free PMC article. Review.
-
Functional Micropeptides Encoded by Long Non-Coding RNAs: A Comprehensive Review.Front Mol Biosci. 2022 Jun 13;9:817517. doi: 10.3389/fmolb.2022.817517. eCollection 2022. Front Mol Biosci. 2022. PMID: 35769907 Free PMC article. Review.
-
Newly Discovered Micropeptide Regulators of SERCA Form Oligomers but Bind to the Pump as Monomers.J Mol Biol. 2019 Nov 8;431(22):4429-4443. doi: 10.1016/j.jmb.2019.07.037. Epub 2019 Aug 23. J Mol Biol. 2019. PMID: 31449798 Free PMC article.
-
The hidden world of membrane microproteins.Exp Cell Res. 2020 Mar 15;388(2):111853. doi: 10.1016/j.yexcr.2020.111853. Epub 2020 Jan 21. Exp Cell Res. 2020. PMID: 31978386 Free PMC article. Review.
-
The role of AMPK in regulation of Na+,K+-ATPase in skeletal muscle: does the gauge always plug the sink?J Muscle Res Cell Motil. 2021 Mar;42(1):77-97. doi: 10.1007/s10974-020-09594-3. Epub 2021 Jan 4. J Muscle Res Cell Motil. 2021. PMID: 33398789 Review.
References
-
- Abascal F, Zardoya R & Posada D (2005). ProtTest: selection of best‐fit models of protein evolution. Bioinformatics 21, 2104–2105. - PubMed
-
- Amores A, Force A, Yan YL, Joly L, Amemiya C, Fritz A, Ho RK, Langeland J, Prince V, Wang YL, Westerfield M, Ekker M & Postlethwait JH (1998). Zebrafish hox clusters and vertebrate genome evolution. Science 282, 1711–1714. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

