The BAF (BRG1/BRM-Associated Factor) chromatin-remodeling complex exhibits ethanol sensitivity in fetal neural progenitor cells and regulates transcription at the miR-9-2 encoding gene locus
- PMID: 28438527
- PMCID: PMC5514786
- DOI: 10.1016/j.alcohol.2017.01.003
The BAF (BRG1/BRM-Associated Factor) chromatin-remodeling complex exhibits ethanol sensitivity in fetal neural progenitor cells and regulates transcription at the miR-9-2 encoding gene locus
Abstract
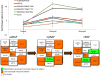
Fetal alcohol spectrum disorders are a leading cause of intellectual disability worldwide. Previous studies have shown that developmental ethanol exposure results in loss of microRNAs (miRNAs), including miR-9, and loss of these miRNAs, in turn, mediates some of ethanol's teratogenic effects in the developing brain. We previously found that ethanol increased methylation at the miR-9-2 encoding gene locus in mouse fetal neural stem cells (NSC), advancing a mechanism for epigenetic silencing of this locus and consequently, miR-9 loss in NSCs. Therefore, we assessed the role of the BAF (BRG1/BRM-Associated Factor) complex, which disassembles nucleosomes to facilitate access to chromatin, as an epigenetic mediator of ethanol's effects on miR-9. Chromatin immunoprecipitation and DNAse I-hypersensitivity analyses showed that the BAF complex was associated with both transcriptionally accessible and heterochromatic regions of the miR-9-2 locus, and that disintegration of the BAF complex by combined knockdown of BAF170 and BAF155 resulted in a significant decrease in miR-9. We hypothesized that ethanol exposure would result in loss of BAF-complex function at the miR-9-2 locus. However, ethanol exposure significantly increased mRNA transcripts for maturation-associated BAF-complex members BAF170, SS18, ARID2, BAF60a, BRM/BAF190b, and BAF53b. Ethanol also significantly increased BAF-complex binding within an intron containing a CpG island and in the terminal exon encoding precursor (pre)-miR-9-2. These data suggest that the BAF complex may adaptively respond to ethanol exposure to protect against a complete loss of miR-9-2 in fetal NSCs. Chromatin remodeling factors may adapt to the presence of a teratogen, to maintain transcription of critical miRNA regulatory pathways.
Keywords: BRG1; DNAse hypersensitivity; Fetal alcohol spectrum disorders; MiR-9.
Copyright © 2017 Elsevier Inc. All rights reserved.
Conflict of interest statement
Conflicts of interest: None
Figures



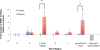

Similar articles
-
A novel Oct4/Pou5f1-like non-coding RNA controls neural maturation and mediates developmental effects of ethanol.Neurotoxicol Teratol. 2021 Jan-Feb;83:106943. doi: 10.1016/j.ntt.2020.106943. Epub 2020 Nov 20. Neurotoxicol Teratol. 2021. PMID: 33221301 Free PMC article.
-
Roles of chromatin remodeling BAF complex in neural differentiation and reprogramming.Cell Tissue Res. 2014 Jun;356(3):575-84. doi: 10.1007/s00441-013-1791-7. Epub 2014 Feb 5. Cell Tissue Res. 2014. PMID: 24496512 Review.
-
Disconnect between alcohol-induced alterations in chromatin structure and gene transcription in a mouse embryonic stem cell model of exposure.Alcohol. 2017 May;60:121-133. doi: 10.1016/j.alcohol.2017.01.007. Epub 2017 Jan 11. Alcohol. 2017. PMID: 28433419 Free PMC article.
-
Dynamic BAF chromatin remodeling complex subunit inclusion promotes temporally distinct gene expression programs in cardiogenesis.Development. 2019 Jul 5;146(19):dev174086. doi: 10.1242/dev.174086. Development. 2019. PMID: 30814119 Free PMC article.
-
Diverse functions of ATP-dependent chromatin remodeling complexes in development and cancer.Acta Biochim Biophys Sin (Shanghai). 2012 Jan;44(1):54-69. doi: 10.1093/abbs/gmr099. Acta Biochim Biophys Sin (Shanghai). 2012. PMID: 22194014 Review.
Cited by
-
Cell-type and fetal-sex-specific targets of prenatal alcohol exposure in developing mouse cerebral cortex.iScience. 2021 Apr 20;24(5):102439. doi: 10.1016/j.isci.2021.102439. eCollection 2021 May 21. iScience. 2021. PMID: 33997709 Free PMC article.
-
miR-9 utilizes precursor pathways in adaptation to alcohol in mouse striatal neurons.Adv Drug Alcohol Res. 2023;3:11323. doi: 10.3389/adar.2023.11323. Epub 2023 Jun 6. Adv Drug Alcohol Res. 2023. PMID: 38116240 Free PMC article.
-
Binge-like ethanol drinking activates anaplastic lymphoma kinase signaling and increases the expression of STAT3 target genes in the mouse hippocampus and prefrontal cortex.Genes Brain Behav. 2021 Feb 28:e12729. doi: 10.1111/gbb.12729. Online ahead of print. Genes Brain Behav. 2021. PMID: 33641239 Free PMC article.
-
Noncoding RNA and Alcohol Use Disorder: A Scoping Review of Current Research and Knowledge Gaps.Alcohol Res. 2025 Jun 20;45(1):06. doi: 10.35946/arcr.v45.1.06. eCollection 2025. Alcohol Res. 2025. PMID: 40552213 Free PMC article.
-
Role of brahma-related gene 1/brahma-associated factor subunits in neural stem/progenitor cells and related neural developmental disorders.World J Stem Cells. 2023 Apr 26;15(4):235-247. doi: 10.4252/wjsc.v15.i4.235. World J Stem Cells. 2023. PMID: 37181007 Free PMC article. Review.
References
-
- Adachi J, Mizoi Y, Fukunaga T, Ogawa Y, Ueno Y, Imamichi H. Degrees of alcohol intoxication in 117 hospitalized cases. J Stud Alcohol. 1991;52(5):448–453. - PubMed
-
- Bultman S, Gebuhr T, Yee D, La Mantia C, Nicholson J, Gilliam A, Randazzo F, Metzger D, Chambon P, Crabtree G, Magnuson T. A Brg1 null mutation in the mouse reveals functional differences among mammalian SWI/SNF complexes. Mol Cell. 2000;6(6):1287–95. - PubMed
-
- Bystron I, Blakemore C, Rakic P. Development of the human cerebral cortex: Boulder Committee revisited. Nat Rev Neurosci. 2008;9(2):110–22. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

