Progress in the Correlative Atomic Force Microscopy and Optical Microscopy
- PMID: 28441775
- PMCID: PMC5426934
- DOI: 10.3390/s17040938
Progress in the Correlative Atomic Force Microscopy and Optical Microscopy
Abstract
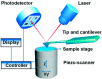
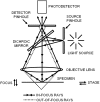
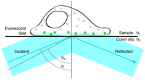
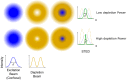
Atomic force microscopy (AFM) has evolved from the originally morphological imaging technique to a powerful and multifunctional technique for manipulating and detecting the interactions between molecules at nanometer resolution. However, AFM cannot provide the precise information of synchronized molecular groups and has many shortcomings in the aspects of determining the mechanism of the interactions and the elaborate structure due to the limitations of the technology, itself, such as non-specificity and low imaging speed. To overcome the technical limitations, it is necessary to combine AFM with other complementary techniques, such as fluorescence microscopy. The combination of several complementary techniques in one instrument has increasingly become a vital approach to investigate the details of the interactions among molecules and molecular dynamics. In this review, we reported the principles of AFM and optical microscopy, such as confocal microscopy and single-molecule localization microscopy, and focused on the development and use of correlative AFM and optical microscopy.
Keywords: atomic force microscopy; conventional florescence microscopy; correlation; super-resolution fluorescence microscopy.
Conflict of interest statement
The authors declare no conflict of interest.
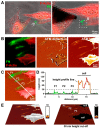
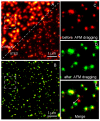
Figures



Similar articles
-
Correlative nanoscopy: A multimodal approach to molecular resolution.Microsc Res Tech. 2021 Oct;84(10):2472-2482. doi: 10.1002/jemt.23800. Epub 2021 May 6. Microsc Res Tech. 2021. PMID: 33955625 Free PMC article. Review.
-
Correlative Super-Resolution Fluorescence Imaging and Atomic Force Microscopy for the Characterization of Biological Samples.Methods Mol Biol. 2017;1663:105-113. doi: 10.1007/978-1-4939-7265-4_9. Methods Mol Biol. 2017. PMID: 28924662
-
How did correlative atomic force microscopy and super-resolution microscopy evolve in the quest for unravelling enigmas in biology?Nanoscale. 2021 Feb 4;13(4):2082-2099. doi: 10.1039/d0nr07203f. Nanoscale. 2021. PMID: 33346312 Review.
-
High-speed near-field fluorescence microscopy combined with high-speed atomic force microscopy for biological studies.Biochim Biophys Acta Gen Subj. 2020 Feb;1864(2):129325. doi: 10.1016/j.bbagen.2019.03.011. Epub 2019 Mar 16. Biochim Biophys Acta Gen Subj. 2020. PMID: 30890438
-
Correlative Super-Resolution Optical and Atomic Force Microscopy Reveals Relationships Between Bacterial Cell Wall Architecture and Synthesis in Bacillus subtilis.ACS Nano. 2021 Oct 26;15(10):16011-16018. doi: 10.1021/acsnano.1c04375. Epub 2021 Sep 17. ACS Nano. 2021. PMID: 34533301 Free PMC article.
Cited by
-
Prior-Apprised Unsupervised Learning of Subpixel Curvilinear Features in Low Signal/Noise Images.Biophys J. 2020 May 19;118(10):2458-2469. doi: 10.1016/j.bpj.2020.04.009. Epub 2020 Apr 19. Biophys J. 2020. PMID: 32359407 Free PMC article.
-
The Contribution of Scanning Force Microscopy on Dental Research: A Narrative Review.Materials (Basel). 2024 Apr 29;17(9):2100. doi: 10.3390/ma17092100. Materials (Basel). 2024. PMID: 38730904 Free PMC article. Review.
-
STORM without enzymatic oxygen scavenging for correlative atomic force and fluorescence superresolution microscopy.Methods Appl Fluoresc. 2018 Jul 9;6(4):045002. doi: 10.1088/2050-6120/aad018. Methods Appl Fluoresc. 2018. PMID: 29956675 Free PMC article.
-
Spatially Resolved Correlation between Stiffness Increase and Actin Aggregation around Nanofibers Internalized in Living Macrophages.Materials (Basel). 2020 Jul 21;13(14):3235. doi: 10.3390/ma13143235. Materials (Basel). 2020. PMID: 32708102 Free PMC article.
-
Cell membrane sample preparation method of combined AFM and dSTORM analysis.Biophys Rep. 2022 Aug 31;8(4):183-192. doi: 10.52601/bpr.2022.220004. Biophys Rep. 2022. PMID: 37288003 Free PMC article.
References
Publication types
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

