Ribonucleotide Reductase Catalytic Subunit M1 (RRM1) as a Novel Therapeutic Target in Multiple Myeloma
- PMID: 28442502
- PMCID: PMC5581671
- DOI: 10.1158/1078-0432.CCR-17-0263
Ribonucleotide Reductase Catalytic Subunit M1 (RRM1) as a Novel Therapeutic Target in Multiple Myeloma
Abstract
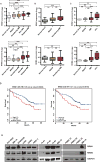
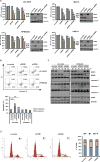
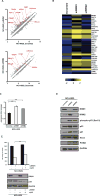
Purpose: To investigate the biological and clinical significance of ribonucleotide reductase (RR) in multiple myeloma.Experimental Design: We assessed the impact of RR expression on patient outcome in multiple myeloma. We then characterized the effect of genetic and pharmacologic inhibition of ribonucleotide reductase catalytic subunit M1 (RRM1) on multiple myeloma growth and survival using siRNA and clofarabine, respectively, in both in vitro and in vivo mouse xenograft models.Results: Newly diagnosed multiple myeloma patients with higher RRM1 expression have shortened survival. Knockdown of RRM1 triggered significant growth inhibition and apoptosis in multiple myeloma cells, even in the context of the bone marrow microenvironment. Gene expression profiling showed upregulation of DNA damage response genes and p53-regulated genes after RRM1 knockdown. Immunoblot and qRT-PCR analysis confirmed that γ-H2A.X, ATM, ATR, Chk1, Chk2, RAD51, 53BP1, BRCA1, and BRCA2 were upregulated/activated. Moreover, immunoblots showed that p53, p21, Noxa, and Puma were activated in p53 wild-type multiple myeloma cells. Clofarabine, a purine nucleoside analogue that inhibits RRM1, induced growth arrest and apoptosis in p53 wild-type cell lines. Although clofarabine did not induce cell death in p53-mutant cells, it did trigger synergistic toxicity in combination with DNA-damaging agent melphalan. Finally, we demonstrated that tumor growth of RRM1-knockdown multiple myeloma cells was significantly reduced in a murine human multiple myeloma cell xenograft model.Conclusions: Our results therefore demonstrate that RRM1 is a novel therapeutic target in multiple myeloma in the preclinical setting and provide the basis for clinical evaluation of RRM1 inhibitor, alone or in combination with DNA-damaging agents, to improve patient outcome in multiple myeloma. Clin Cancer Res; 23(17); 5225-37. ©2017 AACR.
©2017 American Association for Cancer Research.
Conflict of interest statement
Figures





References
-
- Hideshima T, Mitsiades C, Tonon G, Richardson PG, Anderson KC. Understanding multiple myeloma pathogenesis in the bone marrow to identify new therapeutic targets. Nat Rev Cancer. 2007;7:585–98. - PubMed
-
- Lonial S, Dimopoulos M, Palumbo A, White D, Grosicki S, Spicka I, et al. Elotuzumab Therapy for Relapsed or Refractory Multiple Myeloma. N Engl J Med. 2015;373:621–31. - PubMed
-
- Lokhorst HM, Plesner T, Laubach JP, Nahi H, Gimsing P, Hansson M, et al. Targeting CD38 with Daratumumab Monotherapy in Multiple Myeloma. N Engl J Med. 2015;373:1207–19. - PubMed
-
- Watanabe R, Tokuhira M, Kizaki M. Current approaches for the treatment of multiple myeloma. Int J Hematol. 2013;97:333–44. - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Research Materials
Miscellaneous

