GRIM-19 Restricts HCV Replication by Attenuating Intracellular Lipid Accumulation
- PMID: 28443075
- PMCID: PMC5387058
- DOI: 10.3389/fmicb.2017.00576
GRIM-19 Restricts HCV Replication by Attenuating Intracellular Lipid Accumulation
Abstract
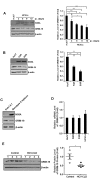
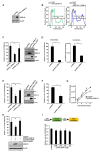
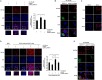
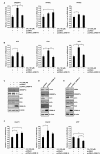
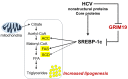
Gene-associated with retinoid-interferon-induced mortality 19 (GRIM-19) targets multiple signaling pathways involved in cell death and growth. However, the role of GRIM-19 in the pathogenesis of hepatitis virus infections remains unexplored. Here, we investigated the restrictive effects of GRIM-19 on the replication of hepatitis C virus (HCV). We found that GRIM-19 protein levels were reduced in HCV-infected Huh7 cells and Huh7 cells harboring HCV replicons. Moreover, ectopically expressed GRIM-19 caused a reduction in both intracellular viral RNA levels and secreted viruses in HCVcc-infected cell cultures. The restrictive effect on HCV replication was restored by treatment with siRNA against GRIM-19. Interestingly, GRIM-19 overexpression did not alter the level of phosphorylated STAT3 or its subcellular distribution. Strikingly, forced expression of GRIM-19 attenuated an increase in intracellular lipid droplets after oleic acid (OA) treatment or HCVcc infection. GRIM-19 overexpression abrogated fatty acid-induced upregulation of sterol regulatory element-binding transcription factor-1 (SREBP-1c), resulting in attenuated expression of its target genes such as fatty acid synthase (FAS) and acetyl CoA carboxylase (ACC). Treatment with OA or overexpression of SREBP-1c in GRIM-19-expressing, HCVcc-infected cells restored HCV replication. Our results suggest that GRIM-19 interferes with HCV replication by attenuating intracellular lipid accumulation and therefore is an anti-viral host factor that could be a promising target for HCV treatment.
Keywords: anti-viral host factor; hepatitis C virus; intracellular lipid accumulation; lipogenesis; viral replication.
Figures


References
-
- Angell J. E., Lindner D. J., Shapiro P. S., Hofmann E. R., Kalvakolanu D. V. (2000). Identification of GRIM-19, a novel cell death-regulatory gene induced by the interferon-beta and retinoic acid combination, using a genetic approach. J. Biol. Chem. 275 33416–33426. 10.1074/jbc.M003929200 - DOI - PubMed
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials
Miscellaneous

