Genotyping by Sequencing for SNP-Based Linkage Analysis and Identification of QTLs Linked to Fruit Quality Traits in Japanese Plum (Prunus salicina Lindl.)
- PMID: 28443103
- PMCID: PMC5386982
- DOI: 10.3389/fpls.2017.00476
Genotyping by Sequencing for SNP-Based Linkage Analysis and Identification of QTLs Linked to Fruit Quality Traits in Japanese Plum (Prunus salicina Lindl.)
Abstract
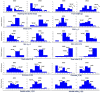
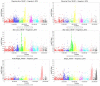
Marker-assisted selection (MAS) in stone fruit (Prunus species) breeding is currently difficult to achieve due to the polygenic nature of the most relevant agronomic traits linked to fruit quality. Genotyping by sequencing (GBS), however, provides a large quantity of useful data suitable for fine mapping using Single Nucleotide Polymorphisms (SNPs) from a reference genome. In this study, GBS was used to genotype 272 seedlings of three F1 Japanese plum (Prunus salicina Lindl) progenies derived from crossing "98-99" (as a common female parent) with "Angeleno," "September King," and "September Queen" as male parents. Raw sequences were aligned to the Peach genome v1, and 42,909 filtered SNPs were obtained after sequence alignment. In addition, 153 seedlings from the "98-99" × "Angeleno" cross were used to develop a genetic map for each parent. A total of 981 SNPs were mapped (479 for "98-99" and 502 for "Angeleno"), covering a genetic distance of 688.8 and 647.03 cM, respectively. Fifty five seedlings from this progeny were phenotyped for different fruit quality traits including ripening time, fruit weight, fruit shape, chlorophyll index, skin color, flesh color, over color, firmness, and soluble solids content in the years 2015 and 2016. Linkage-based QTL analysis allowed the identification of genomic regions significantly associated with ripening time (LG4 of both parents and both phenotyping years), fruit skin color (LG3 and LG4 of both parents and both years), chlorophyll degradation index (LG3 of both parents in 2015) and fruit weight (LG7 of both parents in 2016). These results represent a promising situation for GBS in the identification of SNP variants associated to fruit quality traits, potentially applicable in breeding programs through MAS, in a highly heterozygous crop species such as Japanese plum.
Keywords: GBS; Japanese plum; Prunus salicina; SNP; breeding; fruit quality; molecular markers; ripening.
Figures

Similar articles
-
An Upgraded, Highly Saturated Linkage Map of Japanese Plum (Prunus salicina Lindl.), and Identification of a New Major Locus Controlling the Flavan-3-ol Composition in Fruits.Front Plant Sci. 2022 Mar 4;13:805744. doi: 10.3389/fpls.2022.805744. eCollection 2022. Front Plant Sci. 2022. PMID: 35310655 Free PMC article.
-
Construction of a highly saturated linkage map in Japanese plum (Prunus salicina L.) using GBS for SNP marker calling.PLoS One. 2018 Dec 3;13(12):e0208032. doi: 10.1371/journal.pone.0208032. eCollection 2018. PLoS One. 2018. PMID: 30507961 Free PMC article.
-
De novo transcriptome assembly of 'Angeleno' and 'Lamoon' Japanese plum cultivars (Prunus salicina).Genom Data. 2016 Jun 19;9:35-6. doi: 10.1016/j.gdata.2016.06.010. eCollection 2016 Sep. Genom Data. 2016. PMID: 27408806 Free PMC article.
-
Breeding in peach, cherry and plum: from a tissue culture, genetic, transcriptomic and genomic perspective.Biol Res. 2013;46(3):219-30. doi: 10.4067/S0716-97602013000300001. Biol Res. 2013. PMID: 24346068 Review.
-
Japanese plums (Prunus salicina Lindl.) and phytochemicals--breeding, horticultural practice, postharvest storage, processing and bioactivity.J Sci Food Agric. 2014 Aug;94(11):2137-47. doi: 10.1002/jsfa.6591. Epub 2014 Mar 6. J Sci Food Agric. 2014. PMID: 24449456 Review.
Cited by
-
Adoption and Optimization of Genomic Selection To Sustain Breeding for Apricot Fruit Quality.G3 (Bethesda). 2020 Dec 3;10(12):4513-4529. doi: 10.1534/g3.120.401452. G3 (Bethesda). 2020. PMID: 33067307 Free PMC article.
-
An LTR retrotransposon in the promoter of a PsMYB10.2 gene associated with the regulation of fruit flesh color in Japanese plum.Hortic Res. 2022 Sep 13;9:uhac206. doi: 10.1093/hr/uhac206. eCollection 2022. Hortic Res. 2022. PMID: 36467274 Free PMC article.
-
An Upgraded, Highly Saturated Linkage Map of Japanese Plum (Prunus salicina Lindl.), and Identification of a New Major Locus Controlling the Flavan-3-ol Composition in Fruits.Front Plant Sci. 2022 Mar 4;13:805744. doi: 10.3389/fpls.2022.805744. eCollection 2022. Front Plant Sci. 2022. PMID: 35310655 Free PMC article.
-
Genomic Approaches for Improvement of Tropical Fruits: Fruit Quality, Shelf Life and Nutrient Content.Genes (Basel). 2021 Nov 25;12(12):1881. doi: 10.3390/genes12121881. Genes (Basel). 2021. PMID: 34946829 Free PMC article. Review.
-
Perspectives and recent progress of genome-wide association studies (GWAS) in fruits.Mol Biol Rep. 2022 Jun;49(6):5341-5352. doi: 10.1007/s11033-021-07055-9. Epub 2022 Jan 22. Mol Biol Rep. 2022. PMID: 35064403 Review.
References
-
- Bielenberg D. G., Rauh B., Fan S., Gasic K., Abbott A. G., Reighard G. L., et al. . (2015). Genotyping by sequencing for SNP-based linkage map construction and QTL analysis of chilling requirement and bloom date in peach [Prunus persica (L.) Batsch]. PLoS ONE 10:e0139406. 10.1371/journal.pone.0139406 - DOI - PMC - PubMed
LinkOut - more resources
Full Text Sources
Other Literature Sources

