Quantifying the Number of Independent Organelle DNA Insertions in Genome Evolution and Human Health
- PMID: 28444372
- PMCID: PMC5570036
- DOI: 10.1093/gbe/evx078
Quantifying the Number of Independent Organelle DNA Insertions in Genome Evolution and Human Health
Abstract
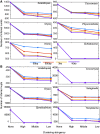
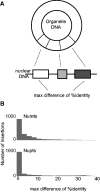
Fragments of organelle genomes are often found as insertions in nuclear DNA. These fragments of mitochondrial DNA (numts) and plastid DNA (nupts) are ubiquitous components of eukaryotic genomes. They are, however, often edited out during the genome assembly process, leading to systematic underestimation of their frequency. Numts and nupts, once inserted, can become further fragmented through subsequent insertion of mobile elements or other recombinational events that disrupt the continuity of the inserted sequence relative to the genuine organelle DNA copy. Because numts and nupts are typically identified through sequence comparison tools such as BLAST, disruption of insertions into smaller fragments can lead to systematic overestimation of numt and nupt frequencies. Accurate identification of numts and nupts is important, however, both for better understanding of their role during evolution, and for monitoring their increasingly evident role in human disease. Human populations are polymorphic for 141 numt loci, five numts are causal to genetic disease, and cancer genomic studies are revealing an abundance of numts associated with tumor progression. Here, we report investigation of salient parameters involved in obtaining accurate estimates of numt and nupt numbers in genome sequence data. Numts and nupts from 44 sequenced eukaryotic genomes reveal lineage-specific differences in the number, relative age and frequency of insertional events as well as lineage-specific dynamics of their postinsertional fragmentation. Our findings outline the main technical parameters influencing accurate identification and frequency estimation of numts in genomic studies pertinent to both evolution and human health.
Keywords: cancer genomics; mitochondria; numts; nupts; organelle insertions.
© The Author 2017. Published by Oxford University Press on behalf of the Society for Molecular Biology and Evolution.
Figures

Similar articles
-
Analysis of plastid and mitochondrial DNA insertions in the nucleus (NUPTs and NUMTs) of six plant species: size, relative age and chromosomal localization.Heredity (Edinb). 2013 Oct;111(4):314-20. doi: 10.1038/hdy.2013.51. Epub 2013 May 29. Heredity (Edinb). 2013. PMID: 23715017 Free PMC article.
-
Mosaic mitochondrial-plastid insertions into the nuclear genome show evidence of both non-homologous end joining and homologous recombination.BMC Evol Biol. 2018 Nov 3;18(1):162. doi: 10.1186/s12862-018-1279-x. BMC Evol Biol. 2018. PMID: 30390623 Free PMC article.
-
NUPTs in sequenced eukaryotes and their genomic organization in relation to NUMTs.Mol Biol Evol. 2004 Oct;21(10):1972-80. doi: 10.1093/molbev/msh210. Epub 2004 Jul 14. Mol Biol Evol. 2004. PMID: 15254258
-
Nuclear Integrants of Organellar DNA Contribute to Genome Structure and Evolution in Plants.Int J Mol Sci. 2020 Jan 21;21(3):707. doi: 10.3390/ijms21030707. Int J Mol Sci. 2020. PMID: 31973163 Free PMC article. Review.
-
Insertions of mitochondrial DNA into the nucleus-effects and role in cell evolution.Genome. 2020 Aug;63(8):365-374. doi: 10.1139/gen-2019-0151. Epub 2020 May 12. Genome. 2020. PMID: 32396758 Review.
Cited by
-
Whole-genome de novo assemblies reveal extensive structural variations and dynamic organelle-to-nucleus DNA transfers in African and Asian rice.Plant J. 2020 Nov;104(3):596-612. doi: 10.1111/tpj.14946. Epub 2020 Aug 27. Plant J. 2020. PMID: 32748498 Free PMC article.
-
The landscape of mitochondrial small non-coding RNAs in the PGCs of male mice, spermatogonia, gametes and in zygotes.BMC Genomics. 2018 Aug 28;19(1):634. doi: 10.1186/s12864-018-5020-3. BMC Genomics. 2018. PMID: 30153810 Free PMC article.
-
A large-scale population based organelle pan-genomes construction and phylogeny analysis reveal the genetic diversity and the evolutionary origins of chloroplast and mitochondrion in Brassica napus L.BMC Genomics. 2022 Apr 30;23(1):339. doi: 10.1186/s12864-022-08573-x. BMC Genomics. 2022. PMID: 35501686 Free PMC article.
-
Two Independent Plastid accD Transfers to the Nuclear Genome of Gnetum and Other Insights on Acetyl-CoA Carboxylase Evolution in Gymnosperms.Genome Biol Evol. 2019 Jun 1;11(6):1691-1705. doi: 10.1093/gbe/evz059. Genome Biol Evol. 2019. PMID: 30924880 Free PMC article.
-
Chromosome-length genome assembly and structural variations of the primal Basenji dog (Canis lupus familiaris) genome.BMC Genomics. 2021 Mar 16;22(1):188. doi: 10.1186/s12864-021-07493-6. BMC Genomics. 2021. PMID: 33726677 Free PMC article.
References
-
- Benesh DP, Hasu T, Suomalainen L-R, Valtonen ET, Tiirola M.. 2006. Reliability of mitochondrial DNA in an acanthocephalan: the problem of pseudogenes. Int J Parasitol. 36:247–254. - PubMed
-
- Bensasson D, Feldman MW, Petrov DA.. 2003. Rates of DNA duplication and mitochondrial DNA insertion in the human genome. J Mol Evol. 57:343–354. - PubMed
-
- Bensasson D, Zhang D, Hartl DL, Hewitt GM.. 2001. Mitochondrial pseudogenes: evolution’s misplaced witnesses. Trends Ecol Evol. 16:314–321. - PubMed
-
- Bertheau C, Schuler H, Ck SK, Arthofer W.. 2011. Hit or miss in phylogeographic analyses: the case of the cryptic NUMTs. Mol Ecol Resour. 11(6):1056–1059. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials

