Structurally Linked Dynamics in Lactate Dehydrogenases of Evolutionarily Distinct Species
- PMID: 28445027
- PMCID: PMC5507060
- DOI: 10.1021/acs.biochem.7b00245
Structurally Linked Dynamics in Lactate Dehydrogenases of Evolutionarily Distinct Species
Abstract
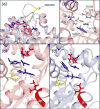
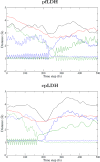
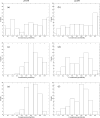
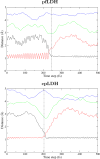
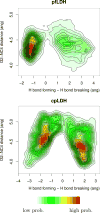
We present new findings about how primary and secondary structure affects the role of fast protein motions in the reaction coordinates of enzymatic reactions. Using transition path sampling and committor distribution analysis, we examined the difference in the role of these fast protein motions in the reaction coordinate of lactate dehydrogenases (LDHs) of Apicomplexa organisms Plasmodium falciparum and Cryptosporidium parvum. Having evolved separately from a common malate dehydrogenase ancestor, the two enzymes exhibit several important structural differences, notably a five-amino acid insertion in the active site loop of P. falciparum LDH. We find that these active site differences between the two organisms' LDHs likely cause a decrease in the contribution of the previously determined LDH rate-promoting vibration to the reaction coordinate of P. falciparum LDH compared to that of C. parvum LDH, specifically in the coupling of the rate-promoting vibration and the hydride transfer. This effect, while subtle, directly shows how changes in structure near the active site of LDH alter catalytically important motions. Insights provided by studying these alterations would prove to be useful in identifying LDH inhibitors that specifically target the isozymes of these parasitic organisms.
Conflict of interest statement
The authors declare no competing financial interest.
Figures
Similar articles
-
Structure, Function, and Thermodynamics of Lactate Dehydrogenases from Humans and the Malaria Parasite P. falciparum.Biochemistry. 2021 Nov 30;60(47):3582-3595. doi: 10.1021/acs.biochem.1c00470. Epub 2021 Nov 8. Biochemistry. 2021. PMID: 34747601 Free PMC article.
-
An atomic-resolution view of neofunctionalization in the evolution of apicomplexan lactate dehydrogenases.Elife. 2014 Jun 25;3:e02304. doi: 10.7554/eLife.02304. Elife. 2014. PMID: 24966208 Free PMC article.
-
Reaction coordinate of an enzymatic reaction revealed by transition path sampling.Proc Natl Acad Sci U S A. 2007 Jul 24;104(30):12253-8. doi: 10.1073/pnas.0704304104. Epub 2007 Jul 17. Proc Natl Acad Sci U S A. 2007. PMID: 17640885 Free PMC article.
-
Lactate dehydrogenase and malate dehydrogenase: Potential antiparasitic targets for drug development studies.Bioorg Med Chem. 2021 Nov 15;50:116458. doi: 10.1016/j.bmc.2021.116458. Epub 2021 Oct 9. Bioorg Med Chem. 2021. PMID: 34687983 Review.
-
Pseudoproteases: mechanisms and function.Biochem J. 2015 May 15;468(1):17-24. doi: 10.1042/BJ20141506. Biochem J. 2015. PMID: 25940733 Review.
Cited by
-
Examining the Origin of Catalytic Power of Catechol O-Methyltransferase.ACS Catal. 2019 Nov 1;9(11):9870-9879. doi: 10.1021/acscatal.9b02657. Epub 2019 Sep 17. ACS Catal. 2019. PMID: 31750009 Free PMC article.
-
Contribution of buried distal amino acid residues in horse liver alcohol dehydrogenase to structure and catalysis.Protein Sci. 2018 Mar;27(3):750-768. doi: 10.1002/pro.3370. Epub 2018 Jan 25. Protein Sci. 2018. PMID: 29271062 Free PMC article.
-
Oscillatory Active-site Motions Correlate with Kinetic Isotope Effects in Formate Dehydrogenase.ACS Catal. 2019 Dec 6;9(12):11199-11206. doi: 10.1021/acscatal.9b03345. Epub 2019 Oct 25. ACS Catal. 2019. PMID: 33996196 Free PMC article.
-
Directed Evolution as a Probe of Rate Promoting Vibrations Introduced via Mutational Change.Biochemistry. 2018 Jun 12;57(23):3289-3298. doi: 10.1021/acs.biochem.8b00185. Epub 2018 Mar 22. Biochemistry. 2018. PMID: 29553716 Free PMC article.
-
Malate dehydrogenase in parasitic protozoans: roles in metabolism and potential therapeutic applications.Essays Biochem. 2024 Oct 3;68(2):235-251. doi: 10.1042/EBC20230075. Essays Biochem. 2024. PMID: 38938216 Free PMC article. Review.
References
-
- Boehr D, McElheny D, Dyson H, Wright P. The dynamic energy landscape of dihydrofolate reductase catalysis. Science. 2006;313:1638–1642. - PubMed
-
- Caratzoulas S, Mincer JS, Schwartz SD. Identification of a protein-promoting vibration in the reaction catalyzed by horse liver alcohol dehydrogenase. J Am Chem Soc. 2002;124:3270–3276. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

