Correlation of histopathologic characteristics to protein expression and function in malignant melanoma
- PMID: 28445515
- PMCID: PMC5405986
- DOI: 10.1371/journal.pone.0176167
Correlation of histopathologic characteristics to protein expression and function in malignant melanoma
Abstract
Background: Metastatic melanoma is still one of the most prevalent skin cancers, which upon progression has neither a prognostic marker nor a specific and lasting treatment. Proteomic analysis is a versatile approach with high throughput data and results that can be used for characterizing tissue samples. However, such analysis is hampered by the complexity of the disease, heterogeneity of patients, tumors, and samples themselves. With the long term aim of quest for better diagnostics biomarkers, as well as predictive and prognostic markers, we focused on relating high resolution proteomics data to careful histopathological evaluation of the tumor samples and patient survival information.
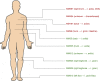
Patients and methods: Regional lymph node metastases obtained from ten patients with metastatic melanoma (stage III) were analyzed by histopathology and proteomics using mass spectrometry. Out of the ten patients, six had clinical follow-up data. The protein deep mining mass spectrometry data was related to the histopathology tumor tissue sections adjacent to the area used for deep-mining. Clinical follow-up data provided information on disease progression which could be linked to protein expression aiming to identify tissue-based specific protein markers for metastatic melanoma and prognostic factors for prediction of progression of stage III disease.
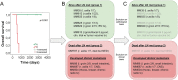
Results: In this feasibility study, several proteins were identified that positively correlated to tumor tissue content including IF6, ARF4, MUC18, UBC12, CSPG4, PCNA, PMEL and MAGD2. The study also identified MYC, HNF4A and TGFB1 as top upstream regulators correlating to tumor tissue content. Other proteins were inversely correlated to tumor tissue content, the most significant being; TENX, EHD2, ZA2G, AOC3, FETUA and THRB. A number of proteins were significantly related to clinical outcome, among these, HEXB, PKM and GPNMB stood out, as hallmarks of processes involved in progression from stage III to stage IV disease and poor survival.
Conclusion: In this feasibility study, promising results show the feasibility of relating proteomics to histopathology and clinical outcome, and insight thus can be gained into the molecular processes driving the disease. The combined analysis of histological features including the sample cellular composition with protein expression of each metastasis enabled the identification of novel, differentially expressed proteins. Further studies are necessary to determine whether these putative biomarkers can be utilized in diagnostics and prognostic prediction of metastatic melanoma.
Conflict of interest statement
Figures


Similar articles
-
Molecular oncogene markers and their significance in cutaneous malignant melanoma.Ann Surg Oncol. 1998 Apr-May;5(3):253-60. doi: 10.1007/BF02303782. Ann Surg Oncol. 1998. PMID: 9607628
-
Heterogenous S-100B protein expression patterns in malignant melanoma and association with serum protein levels.Oncology. 2003;64(4):374-9. doi: 10.1159/000070296. Oncology. 2003. PMID: 12759535
-
High expression of glycolytic and pigment proteins is associated with worse clinical outcome in stage III melanoma.Melanoma Res. 2013 Dec;23(6):452-60. doi: 10.1097/CMR.0000000000000027. Melanoma Res. 2013. PMID: 24128789
-
Melanoma patient staging: histopathological versus molecular evaluation of the sentinel node.Melanoma Res. 2003 Jun;13(3):313-24. doi: 10.1097/01.cmr.0000056229.78713.7d. Melanoma Res. 2003. PMID: 12777989 Review.
-
Evidence and interdisciplinary consense-based German guidelines: diagnosis and surveillance of melanoma.Melanoma Res. 2007 Dec;17(6):393-9. doi: 10.1097/CMR.0b013e3282f05039. Melanoma Res. 2007. PMID: 17992123
Cited by
-
Challenging the heterogeneity of disease presentation in malignant melanoma-impact on patient treatment.Cell Biol Toxicol. 2019 Feb;35(1):1-14. doi: 10.1007/s10565-018-9446-9. Epub 2018 Oct 24. Cell Biol Toxicol. 2019. PMID: 30357519 Free PMC article. Review.
-
The Hidden Story of Heterogeneous B-raf V600E Mutation Quantitative Protein Expression in Metastatic Melanoma-Association with Clinical Outcome and Tumor Phenotypes.Cancers (Basel). 2019 Dec 9;11(12):1981. doi: 10.3390/cancers11121981. Cancers (Basel). 2019. PMID: 31835364 Free PMC article.
-
IL-6-Driven Autocrine Lactate Promotes Immune Escape of Uveal Melanoma.Invest Ophthalmol Vis Sci. 2024 Mar 5;65(3):37. doi: 10.1167/iovs.65.3.37. Invest Ophthalmol Vis Sci. 2024. PMID: 38551584 Free PMC article.
-
Improved survival prognostication of node-positive malignant melanoma patients utilizing shotgun proteomics guided by histopathological characterization and genomic data.Sci Rep. 2019 Mar 26;9(1):5154. doi: 10.1038/s41598-019-41625-z. Sci Rep. 2019. PMID: 30914758 Free PMC article.
-
Hallmarks of glycogene expression and glycosylation pathways in squamous and adenocarcinoma cervical cancer.PeerJ. 2021 Aug 31;9:e12081. doi: 10.7717/peerj.12081. eCollection 2021. PeerJ. 2021. PMID: 34540372 Free PMC article.
References
-
- Thiam A, Zhao Z, Quinn C, Barber B. Years of life lost due to metastatic melanoma in 12 countries. Journal of medical economics. 2016;19(3):259–64. Epub 2015/11/05. doi: 10.3111/13696998.2015.1115764 - DOI - PubMed
-
- Berwick M, Buller DB, Cust A, Gallagher R, Lee TK, Meyskens F, et al. Melanoma Epidemiology and Prevention. Cancer treatment and research. 2016;167:17–49. Epub 2015/11/26. doi: 10.1007/978-3-319-22539-5_2 - DOI - PubMed
-
- Eriksson H, Lyth J, Andersson TM. The proportion cured of patients diagnosed with Stage III-IV cutaneous malignant melanoma in Sweden 1990–2007: A population-based study. International journal of cancer Journal international du cancer. 2016;138(12):2829–36. Epub 2016/01/28. doi: 10.1002/ijc.30023 - DOI - PubMed
-
- Hanahan D, Weinberg RA. The hallmarks of cancer. Cell. 2000;100(1):57–70. Epub 2000/01/27. - PubMed
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Miscellaneous

