Bim gene dosage is critical in modulating nephron progenitor survival in the absence of microRNAs during kidney development
- PMID: 28446592
- PMCID: PMC5503708
- DOI: 10.1096/fj.201700010R
Bim gene dosage is critical in modulating nephron progenitor survival in the absence of microRNAs during kidney development
Abstract
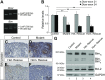
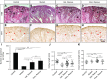
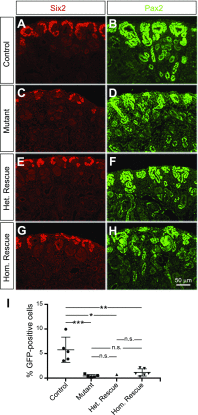
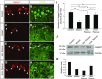
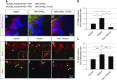
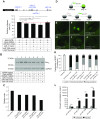
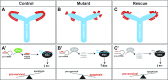
Low nephron endowment at birth has been associated with an increased risk for developing hypertension and chronic kidney disease. We demonstrated in an earlier study that conditional deletion of the microRNA (miRNA)-processing enzyme Dicer from nephron progenitors results in premature depletion of the progenitors and increased expression of the proapoptotic protein Bim (also known as Bcl-2L11). In this study, we generated a compound mouse model with conditional deletion of both Dicer and Bim, to determine the biologic significance of increased Bim expression in Dicer-deficient nephron progenitors. The loss of Bim partially restored the number of nephron progenitors and improved nephron formation. The number of progenitors undergoing apoptosis was significantly reduced in kidneys with loss of a single allele, or both alleles, of Bim compared to mutant kidneys. Furthermore, 2 miRNAs expressed in nephron progenitors (miR-17 and miR-106b) regulated Bim levels in vitro and in vivo Together, these data suggest that miRNA-mediated regulation of Bim controls nephron progenitor survival during nephrogenesis, as one potential means of regulating nephron endowment.-Cerqueira, D. M., Bodnar, A. J., Phua, Y. L., Freer, R., Hemker, S. L., Walensky, L. D., Hukriede, N. A., Ho, J. Bim gene dosage is critical in modulating nephron progenitor survival in the absence of microRNAs during kidney development.
Keywords: Bcl-2L11; Dicer; apoptosis; metanephros; nephrogenesis.
© FASEB.
Figures
References
-
- Vize P. D., Woolf A. S., Bard J. (2003) The Kidney: From Normal Development to Congenital Disease, Academic, Amsterdam/London
-
- Lechner M. S., Dressler G. R. (1997) The molecular basis of embryonic kidney development. Mech. Dev. 62, 105–120 - PubMed
-
- Saxén L., Sariola H. (1987) Early organogenesis of the kidney. Pediatr. Nephrol. 1, 385–392 - PubMed
-
- Hinchliffe S. A., Sargent P. H., Howard C. V., Chan Y. F., van Velzen D. (1991) Human intrauterine renal growth expressed in absolute number of glomeruli assessed by the disector method and Cavalieri principle. Lab. Invest. 64, 777–784 - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases

