The histone variant H2A.Z promotes splicing of weak introns
- PMID: 28446597
- PMCID: PMC5411709
- DOI: 10.1101/gad.295287.116
The histone variant H2A.Z promotes splicing of weak introns
Abstract
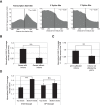
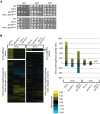
Multiple lines of evidence implicate chromatin in the regulation of premessenger RNA (pre-mRNA) splicing. However, the influence of chromatin factors on cotranscriptional splice site usage remains unclear. Here we investigated the function of the highly conserved histone variant H2A.Z in pre-mRNA splicing using the intron-rich model yeast Schizosaccharomyces pombe Using epistatic miniarray profiles (EMAPs) to survey the genetic interaction landscape of the Swr1 nucleosome remodeling complex, which deposits H2A.Z, we uncovered evidence for functional interactions with components of the spliceosome. In support of these genetic connections, splicing-specific microarrays show that H2A.Z and the Swr1 ATPase are required during temperature stress for the efficient splicing of a subset of introns. Notably, affected introns are enriched for H2A.Z occupancy and more likely to contain nonconsensus splice sites. To test the significance of the latter correlation, we mutated the splice sites in an affected intron to consensus and found that this suppressed the requirement for H2A.Z in splicing of that intron. These data suggest that H2A.Z occupancy promotes cotranscriptional splicing of suboptimal introns that may otherwise be discarded via proofreading ATPases. Consistent with this model, we show that overexpression of splicing ATPase Prp16 suppresses both the growth and splicing defects seen in the absence of H2A.Z.
Keywords: H2A.Z; Prp16; Swr1 complex; chromatin; fission yeast; pre-mRNA splicing.
© 2017 Nissen et al.; Published by Cold Spring Harbor Laboratory Press.
Figures





References
-
- Ast G. 2004. How did alternative splicing evolve? Nat Rev Genet 5: 773–782. - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
