plantiSMASH: automated identification, annotation and expression analysis of plant biosynthetic gene clusters
- PMID: 28453650
- PMCID: PMC5570173
- DOI: 10.1093/nar/gkx305
plantiSMASH: automated identification, annotation and expression analysis of plant biosynthetic gene clusters
Abstract
Plant specialized metabolites are chemically highly diverse, play key roles in host-microbe interactions, have important nutritional value in crops and are frequently applied as medicines. It has recently become clear that plant biosynthetic pathway-encoding genes are sometimes densely clustered in specific genomic loci: biosynthetic gene clusters (BGCs). Here, we introduce plantiSMASH, a versatile online analysis platform that automates the identification of candidate plant BGCs. Moreover, it allows integration of transcriptomic data to prioritize candidate BGCs based on the coexpression patterns of predicted biosynthetic enzyme-coding genes, and facilitates comparative genomic analysis to study the evolutionary conservation of each cluster. Applied on 48 high-quality plant genomes, plantiSMASH identifies a rich diversity of candidate plant BGCs. These results will guide further experimental exploration of the nature and dynamics of gene clustering in plant metabolism. Moreover, spurred by the continuing decrease in costs of plant genome sequencing, they will allow genome mining technologies to be applied to plant natural product discovery. The plantiSMASH web server, precalculated results and source code are freely available from http://plantismash.secondarymetabolites.org.
© The Author(s) 2017. Published by Oxford University Press on behalf of Nucleic Acids Research.
Figures
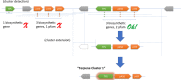



References
-
- Rutledge P.J., Challis G.L.. Discovery of microbial natural products by activation of silent biosynthetic gene clusters. Nat. Rev. Microbiol. 2015; 13:509–523. - PubMed
-
- Ziemert N., Alanjary M., Weber T.. The evolution of genome mining in microbes - a review. Nat. Prod. Rep. 2016; 33:988–1005. - PubMed
-
- Medema M.H., Blin K., Cimermancic P., de Jager V., Zakrzewski P., Fischbach M.A., Weber T., Takano E., Breitling R.. antiSMASH: rapid identification, annotation and analysis of secondary metabolite biosynthesis gene clusters in bacterial and fungal genome sequences. Nucleic Acids Res. 2011; 39:W339–W346. - PMC - PubMed
Publication types
MeSH terms
Substances
Grants and funding
- BBS/E/J/000PR9790/BB_/Biotechnology and Biological Sciences Research Council/United Kingdom
- BBS/E/J/00000614/BB_/Biotechnology and Biological Sciences Research Council/United Kingdom
- BB/L014130/1/BB_/Biotechnology and Biological Sciences Research Council/United Kingdom
- BB/M028860/1/BB_/Biotechnology and Biological Sciences Research Council/United Kingdom
- U01 GM110699/GM/NIGMS NIH HHS/United States
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

