Quantitative identification of senescent cells in aging and disease
- PMID: 28455874
- PMCID: PMC5506427
- DOI: 10.1111/acel.12592
Quantitative identification of senescent cells in aging and disease
Abstract
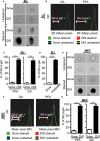
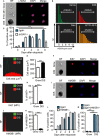
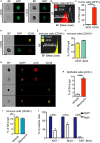
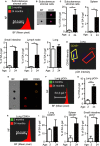
Senescent cells are present in premalignant lesions and sites of tissue damage and accumulate in tissues with age. In vivo identification, quantification and characterization of senescent cells are challenging tasks that limit our understanding of the role of senescent cells in diseases and aging. Here, we present a new way to precisely quantify and identify senescent cells in tissues on a single-cell basis. The method combines a senescence-associated beta-galactosidase assay with staining of molecular markers for cellular senescence and of cellular identity. By utilizing technology that combines flow cytometry with high-content image analysis, we were able to quantify senescent cells in tumors, fibrotic tissues, and tissues of aged mice. Our approach also yielded the finding that senescent cells in tissues of aged mice are larger than nonsenescent cells. Thus, this method provides a basis for quantitative assessment of senescent cells and it offers proof of principle for combination of different markers of senescence. It paves the way for screening of senescent cells for identification of new senescence biomarkers, genes that bypass senescence or senolytic compounds that eliminate senescent cells, thus enabling a deeper understanding of the senescent state in vivo.
Keywords: ImageStreamX; aging; cancer; cellular senescence; flow cytometry; senescence-associated beta-galactosidase.
© 2017 The Authors. Aging Cell published by the Anatomical Society and John Wiley & Sons Ltd.
Figures

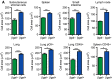
References
-
- Aoshiba K, Tsuji T, Kameyama S, Itoh M, Semba S, Yamaguchi K, Nakamura H (2013) Senescence‐associated secretory phenotype in a mouse model of bleomycin‐induced lung injury. Exp. Toxicol. Pathol. 65, 1053–1062. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical

