Gene Expression Signature Differentiates Histology But Not Progression Status of Early-Stage NSCLC
- PMID: 28456114
- PMCID: PMC5408153
- DOI: 10.1016/j.tranon.2017.01.015
Gene Expression Signature Differentiates Histology But Not Progression Status of Early-Stage NSCLC
Abstract
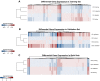
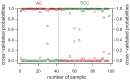
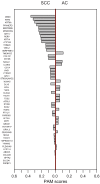
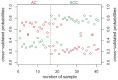
Advances in molecular analyses based on high-throughput technologies can contribute to a more accurate classification of non-small cell lung cancer (NSCLC), as well as a better prediction of both the disease course and the efficacy of targeted therapies. Here we set out to analyze whether global gene expression profiling performed in a group of early-stage NSCLC patients can contribute to classifying tumor subtypes and predicting the disease prognosis. Gene expression profiling was performed with the use of the microarray technology in a training set of 108 NSCLC samples. Subsequently, the recorded findings were validated further in an independent cohort of 44 samples. We demonstrated that the specific gene patterns differed significantly between lung adenocarcinoma (AC) and squamous cell lung carcinoma (SCC) samples. Furthermore, we developed and validated a novel 53-gene signature distinguishing SCC from AC with 93% accuracy. Evaluation of the classifier performance in the validation set showed that our predictor classified the AC patients with 100% sensitivity and 88% specificity. We revealed that gene expression patterns observed in the early stages of NSCLC may help elucidate the histological distinctions of tumors through identification of different gene-mediated biological processes involved in the pathogenesis of histologically distinct tumors. However, we showed here that the gene expression profiles did not provide additional value in predicting the progression status of the early-stage NSCLC. Nevertheless, the gene expression signature analysis enabled us to perform a reliable subclassification of NSCLC tumors, and it can therefore become a useful diagnostic tool for a more accurate selection of patients for targeted therapies.
Copyright © 2017 The Authors. Published by Elsevier Inc. All rights reserved.
Figures
Similar articles
-
Diagnostic assay based on hsa-miR-205 expression distinguishes squamous from nonsquamous non-small-cell lung carcinoma.J Clin Oncol. 2009 Apr 20;27(12):2030-7. doi: 10.1200/JCO.2008.19.4134. Epub 2009 Mar 9. J Clin Oncol. 2009. PMID: 19273703
-
Validation for histology-driven diagnosis in non-small cell lung cancer using hsa-miR-205 and hsa-miR-21 expression by two different normalization strategies.Int J Cancer. 2016 Feb 1;138(3):689-97. doi: 10.1002/ijc.29816. Epub 2015 Sep 10. Int J Cancer. 2016. PMID: 26311306
-
Classification of Non-Small Cell Lung Cancer Using Significance Analysis of Microarray-Gene Set Reduction Algorithm.Biomed Res Int. 2016;2016:2491671. doi: 10.1155/2016/2491671. Epub 2016 Jun 30. Biomed Res Int. 2016. PMID: 27446945 Free PMC article.
-
[Non-small cell lung cancer. New biomarkers for diagnostics and therapy].Pathologe. 2015 Nov;36 Suppl 2:189-93. doi: 10.1007/s00292-015-0084-1. Pathologe. 2015. PMID: 26391246 Review. German.
-
Are adenosquamous lung carcinomas a simple mix of adenocarcinomas and squamous cell carcinomas, or more complex at the molecular level?Lung Cancer. 2010 Apr;68(1):1-9. doi: 10.1016/j.lungcan.2009.11.001. Epub 2009 Dec 8. Lung Cancer. 2010. PMID: 20004040 Review.
Cited by
-
Novel blood-based FUT7 DNA methylation is associated with lung cancer: especially for lung squamous cell carcinoma.Clin Epigenetics. 2022 Dec 3;14(1):167. doi: 10.1186/s13148-022-01389-2. Clin Epigenetics. 2022. PMID: 36463240 Free PMC article.
-
Tumor-based gene expression biomarkers to predict survival following curative intent resection for stage I lung adenocarcinoma.PLoS One. 2018 Nov 20;13(11):e0207513. doi: 10.1371/journal.pone.0207513. eCollection 2018. PLoS One. 2018. PMID: 30458017 Free PMC article.
-
Serum Insights: Leveraging the Power of miRNA Profiling as an Early Diagnostic Tool for Non-Small Cell Lung Cancer.Cancers (Basel). 2023 Oct 10;15(20):4910. doi: 10.3390/cancers15204910. Cancers (Basel). 2023. PMID: 37894277 Free PMC article.
-
miRNA-Seq Tissue Diagnostic Signature: A Novel Model for NSCLC Subtyping.Int J Mol Sci. 2023 Aug 28;24(17):13318. doi: 10.3390/ijms241713318. Int J Mol Sci. 2023. PMID: 37686123 Free PMC article.
-
Identification of Tumor Microenvironment-Related Prognostic lncRNAs in Lung Adenocarcinoma.Front Oncol. 2021 Aug 2;11:719812. doi: 10.3389/fonc.2021.719812. eCollection 2021. Front Oncol. 2021. PMID: 34408984 Free PMC article.
References
-
- Torre LA, Siegel RL, Jemal A. Lung cancer statistics. Adv Exp Med Biol. 2016;893:1–19. - PubMed
-
- Manegold C. Treatment algorithm in 2014 for advanced non–small cell lung cancer: therapy selection by tumour histology and molecular biology. Adv Med Sci. 2014;59:308–313. - PubMed
-
- Jorda M, Gomez-Fernandez C, Garcia M, Mousavi F, Walker G, Mejias A, Fernandez-Castro G, Ganjei-Azar P. P63 differentiates subtypes of nonsmall cell carcinomas of lung in cytologic samples: implications in treatment selection. Cancer Cytopathol. 2009;117:46–50. - PubMed
-
- Stang A, Pohlabeln H, Muller KM, Jahn I, Giersiepen K, Jockel KH. Diagnostic agreement in the histopathological evaluation of lung cancer tissue in a population-based case-control study. Lung Cancer. 2006;52:29–36. - PubMed
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials

