Towards precision prevention: Technologies for identifying healthy individuals with high risk of disease
- PMID: 28458064
- PMCID: PMC5841554
- DOI: 10.1016/j.mrfmmm.2017.03.007
Towards precision prevention: Technologies for identifying healthy individuals with high risk of disease
Abstract
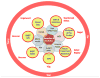
The rise of advanced technologies for characterizing human populations at the molecular level, from sequence to function, is shifting disease prevention paradigms toward personalized strategies. Because minimization of adverse outcomes is a key driver for treatment decisions for diseased populations, developing personalized therapy strategies represent an important dimension of both precision medicine and personalized prevention. In this commentary, we highlight recently developed enabling technologies in the field of DNA damage, DNA repair, and mutagenesis. We propose that omics approaches and functional assays can be integrated into population studies that fuse basic, translational and clinical research with commercial expertise in order to accelerate personalized prevention and treatment of cancer and other diseases linked to aberrant responses to DNA damage. This collaborative approach is generally applicable to efforts to develop data-driven, individualized prevention and treatment strategies for other diseases. We also recommend strategies for maximizing the use of biological samples for epidemiological studies, and for applying emerging technologies to clinical applications.
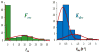
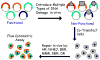
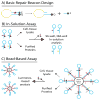
Keywords: Comet; DNA damage; DNA damage response; DNA repair; H2AX; Host cell reactivation; Precision medicine.
Copyright © 2017 Elsevier B.V. All rights reserved.
Conflict of interest statement
J.H.B. has consultancy, stock ownership, and royalties with Grail Inc. B.P.E. is a co-inventor on a patent for CometChip that has been licensed to Trevigen, Inc. R.W.S. is a scientific consultant for Trevigen, Inc.
Figures
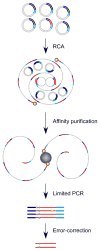
References
-
- Auerbach C. Chemical Mutagenesis. Biol Rev Camb Philos Soc. 1949;24:355–391. - PubMed
-
- HoIlaender A, Emmons CW. Wavelength dependence of mutation production in the ultraviolet with special emphasis on fungi. Cold Spring Harb Symp Quant Biol. 1941:179–186.
-
- Hollaender A, Baker WK, Anderson EH. Effect of oxygen tension and certain chemicals on the x-ray sensitivity of mutation production and survival. Cold Spring Harb Symp Quant Biol. 1951;16:315–326. - PubMed
-
- Weigle JJ, Bertani G. Multiplicity reactivation of bacteriophage inactivated by ionizing radiations. Virology. 1956;2:344–355. - PubMed
Publication types
MeSH terms
Grants and funding
- U01 ES011038/ES/NIEHS NIH HHS/United States
- R21 ES019492/ES/NIEHS NIH HHS/United States
- R01 ES011740/ES/NIEHS NIH HHS/United States
- Z99 ES999999/ImNIH/Intramural NIH HHS/United States
- P01 CA092584/CA/NCI NIH HHS/United States
- DP1 ES022576/ES/NIEHS NIH HHS/United States
- R44 ES021116/ES/NIEHS NIH HHS/United States
- R01 ES019319/ES/NIEHS NIH HHS/United States
- R44 ES025139/ES/NIEHS NIH HHS/United States
- R01 CA148629/CA/NCI NIH HHS/United States
- R21 ES019494/ES/NIEHS NIH HHS/United States
- P30 ES009089/ES/NIEHS NIH HHS/United States
LinkOut - more resources
Full Text Sources
Other Literature Sources

