In vivo mutation rates and the landscape of fitness costs of HIV-1
- PMID: 28458914
- PMCID: PMC5399928
- DOI: 10.1093/ve/vex003
In vivo mutation rates and the landscape of fitness costs of HIV-1
Abstract
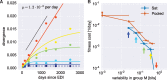
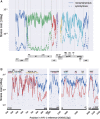
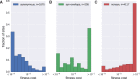
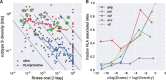
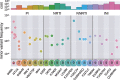
Mutation rates and fitness costs of deleterious mutations are difficult to measure in vivo but essential for a quantitative understanding of evolution. Using whole genome deep sequencing data from longitudinal samples during untreated HIV-1 infection, we estimated mutation rates and fitness costs in HIV-1 from the dynamics of genetic variation. At approximately neutral sites, mutations accumulate with a rate of 1.2 × 10-5 per site per day, in agreement with the rate measured in cell cultures. We estimated the rate from G to A to be the largest, followed by the other transitions C to T, T to C, and A to G, while transversions are less frequent. At other sites, mutations tend to reduce virus replication. We estimated the fitness cost of mutations at every site in the HIV-1 genome using a model of mutation selection balance. About half of all non-synonymous mutations have large fitness costs (>10 percent), while most synonymous mutations have costs <1 percent. The cost of synonymous mutations is especially low in most of pol where we could not detect measurable costs for the majority of synonymous mutations. In contrast, we find high costs for synonymous mutations in important RNA structures and regulatory regions. The intra-patient fitness cost estimates are consistent across multiple patients, indicating that the deleterious part of the fitness landscape is universal and explains a large fraction of global HIV-1 group M diversity.
Keywords: Evolution; HIV-1; fitness landscape.; mutation rate.
Figures

References
-
- Abbotts J., Bebenek K., Kunkel T. A., Wilson S. H. (1993) ‘Termination of processive synthesis on a natural DNA template is influenced by the sequence of the template-primer stem’, Journal of Biological Chemistry, 268: 10312.. - PubMed
LinkOut - more resources
Full Text Sources
Other Literature Sources

