Functional screening of aldehyde decarbonylases for long-chain alkane production by Saccharomyces cerevisiae
- PMID: 28464872
- PMCID: PMC5414326
- DOI: 10.1186/s12934-017-0683-z
Functional screening of aldehyde decarbonylases for long-chain alkane production by Saccharomyces cerevisiae
Abstract
Background: Low catalytic activities of pathway enzymes are often a limitation when using microbial based chemical production. Recent studies indicated that the enzyme activity of aldehyde decarbonylase (AD) is a critical bottleneck for alkane biosynthesis in Saccharomyces cerevisiae. We therefore performed functional screening to identify efficient ADs that can improve alkane production by S. cerevisiae.
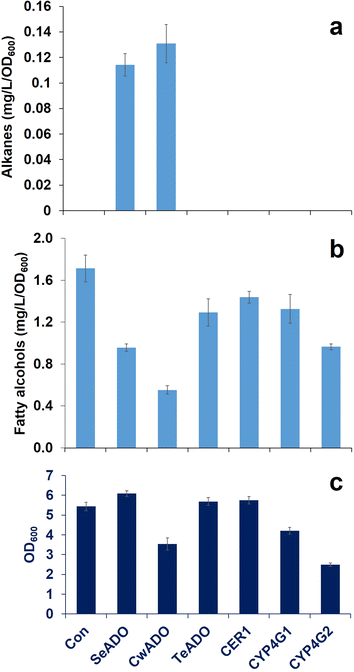
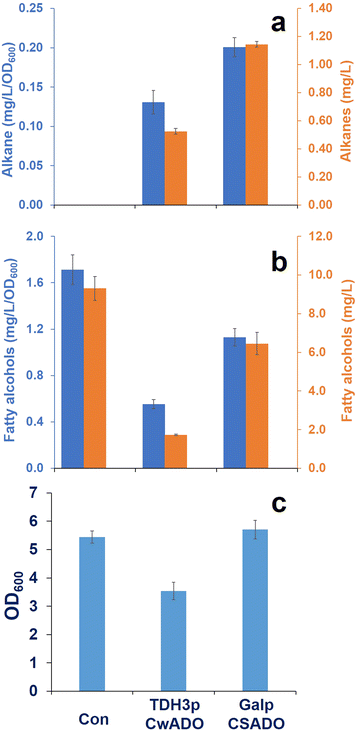
Results: A comparative study of ADs originated from a plant, insects, and cyanobacteria were conducted in S. cerevisiae. As a result, expression of aldehyde deformylating oxygenases (ADOs), which are cyanobacterial ADs, from Synechococcus elongatus and Crocosphaera watsonii converted fatty aldehydes to corresponding Cn-1 alkanes and alkenes. The CwADO showed the highest alkane titer (0.13 mg/L/OD600) and the lowest fatty alcohol production (0.55 mg/L/OD600). However, no measurable alkanes and alkenes were detected in other AD expressed yeast strains. Dynamic expression of SeADO and CwADO under GAL promoters increased alkane production to 0.20 mg/L/OD600 and no fatty alcohols, with even number chain lengths from C8 to C14, were detected in the cells.
Conclusions: We demonstrated in vivo enzyme activities of ADs by displaying profiles of alkanes and fatty alcohols in S. cerevisiae. Among the AD enzymes evaluated, cyanobacteria ADOs were found to be suitable for alkane biosynthesis in S. cerevisiae. This work will be helpful to decide an AD candidate for alkane biosynthesis in S. cerevisiae and it will provide useful information for further investigation of AD enzymes with improved activities.
Keywords: Aldehyde decarbonylase; Alkane biosynthesis; Biofuels; Metabolic engineering; Saccharomyces cerevisiae.
Figures

Similar articles
-
Microbial biosynthesis of alkanes.Science. 2010 Jul 30;329(5991):559-62. doi: 10.1126/science.1187936. Science. 2010. PMID: 20671186
-
Utilizing Alcohol for Alkane Biosynthesis by Introducing a Fatty Alcohol Dehydrogenase.Appl Environ Microbiol. 2022 Dec 13;88(23):e0126422. doi: 10.1128/aem.01264-22. Epub 2022 Nov 23. Appl Environ Microbiol. 2022. PMID: 36416567 Free PMC article.
-
Enabling the synthesis of medium chain alkanes and 1-alkenes in yeast.Metab Eng. 2017 Nov;44:81-88. doi: 10.1016/j.ymben.2017.09.007. Epub 2017 Sep 20. Metab Eng. 2017. PMID: 28939277
-
Cyanobacterial Enzymes for Bioalkane Production.Adv Exp Med Biol. 2018;1080:119-154. doi: 10.1007/978-981-13-0854-3_6. Adv Exp Med Biol. 2018. PMID: 30091094 Review.
-
Recent advances in the improvement of cyanobacterial enzymes for bioalkane production.Microb Cell Fact. 2022 Dec 12;21(1):256. doi: 10.1186/s12934-022-01981-4. Microb Cell Fact. 2022. PMID: 36503511 Free PMC article. Review.
Cited by
-
Insights into cyanobacterial alkane biosynthesis.J Ind Microbiol Biotechnol. 2022 Apr 14;49(2):kuab075. doi: 10.1093/jimb/kuab075. J Ind Microbiol Biotechnol. 2022. PMID: 34718648 Free PMC article. Review.
-
Drop-in biofuel production using fatty acid photodecarboxylase from Chlorella variabilis in the oleaginous yeast Yarrowia lipolytica.Biotechnol Biofuels. 2019 Aug 24;12:202. doi: 10.1186/s13068-019-1542-4. eCollection 2019. Biotechnol Biofuels. 2019. PMID: 31462926 Free PMC article.
-
The Prokaryotic Microalga Limnothrix redekei KNUA012 to Improve Aldehyde Decarbonylase Expression for Use as a Biological Resource.Pol J Microbiol. 2023 Sep 20;72(3):307-317. doi: 10.33073/pjm-2023-031. eCollection 2023 Sep 1. Pol J Microbiol. 2023. PMID: 37725893 Free PMC article.
-
Advanced metabolic Engineering strategies for the sustainable production of free fatty acids and their derivatives using yeast.J Biol Eng. 2024 Dec 27;18(1):73. doi: 10.1186/s13036-024-00473-w. J Biol Eng. 2024. PMID: 39731138 Free PMC article. Review.
-
Engineering Saccharomyces cerevisiae cells for production of fatty acid-derived biofuels and chemicals.Open Biol. 2019 May 31;9(5):190049. doi: 10.1098/rsob.190049. Open Biol. 2019. PMID: 31088249 Free PMC article. Review.
References
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials
Miscellaneous

