Illumina MiSeq 16S amplicon sequence analysis of bovine respiratory disease associated bacteria in lung and mediastinal lymph node tissue
- PMID: 28464950
- PMCID: PMC5414144
- DOI: 10.1186/s12917-017-1035-2
Illumina MiSeq 16S amplicon sequence analysis of bovine respiratory disease associated bacteria in lung and mediastinal lymph node tissue
Abstract
Background: Bovine respiratory disease (BRD) is caused by growth of single or multiple species of pathogenic bacteria in lung tissue following stress and/or viral infection. Next generation sequencing of 16S ribosomal RNA gene PCR amplicons (NGS 16S amplicon analysis) is a powerful culture-independent open reference method that has recently been used to increase understanding of BRD-associated bacteria in the upper respiratory tract of BRD cattle. However, it has not yet been used to examine the microbiome of the bovine lower respiratory tract. The objective of this study was to use NGS 16S amplicon analysis to identify bacteria in post-mortem lung and lymph node tissue samples harvested from fatal BRD cases and clinically healthy animals. Cranial lobe and corresponding mediastinal lymph node post-mortem tissue samples were collected from calves diagnosed as BRD cases by veterinary laboratory pathologists and from clinically healthy calves. NGS 16S amplicon libraries, targeting the V3-V4 region of the bacterial 16S rRNA gene were prepared and sequenced on an Illumina MiSeq. Quantitative insights into microbial ecology (QIIME) was used to determine operational taxonomic units (OTUs) which corresponded to the 16S rRNA gene sequences.
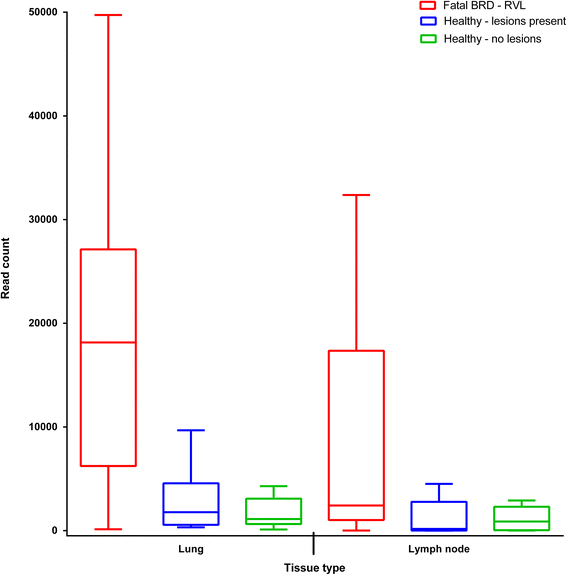
Results: Leptotrichiaceae, Mycoplasma, Pasteurellaceae, and Fusobacterium were the most abundant OTUs identified in the lungs and lymph nodes of the calves which died from BRD. Leptotrichiaceae, Fusobacterium, Mycoplasma, Trueperella and Bacteroides had greater relative abundances in post-mortem lung samples collected from fatal cases of BRD in dairy calves, compared with clinically healthy calves without lung lesions. Leptotrichiaceae, Mycoplasma and Pasteurellaceae showed higher relative abundances in post-mortem lymph node samples collected from fatal cases of BRD in dairy calves, compared with clinically healthy calves without lung lesions. Two Leptotrichiaceae sequence contigs were subsequently assembled from bacterial DNA-enriched shotgun sequences.
Conclusions: The microbiomes of the cranial lung lobe and mediastinal lymph node from calves which died from BRD and from clinically healthy H-F calves have been characterised. Contigs corresponding to the abundant Leptotrichiaceae OTU were sequenced and found not to be identical to any known bacterial genus. This suggests that we have identified a novel bacterial species associated with BRD.
Keywords: 16S sequencing; Bovine respiratory disease; diagnostics; lung microbiome.
Figures


Similar articles
-
Disparity in the nasopharyngeal microbiota between healthy cattle on feed, at entry processing and with respiratory disease.Vet Microbiol. 2017 Sep;208:30-37. doi: 10.1016/j.vetmic.2017.07.006. Epub 2017 Jul 13. Vet Microbiol. 2017. PMID: 28888646
-
Characterization of the upper and lower respiratory tract microbiota in Piedmontese calves.Microbiome. 2017 Nov 21;5(1):152. doi: 10.1186/s40168-017-0372-5. Microbiome. 2017. PMID: 29157308 Free PMC article.
-
Comparison of the nasopharyngeal bacterial microbiota of beef calves raised without the use of antimicrobials between healthy calves and those diagnosed with bovine respiratory disease.Vet Microbiol. 2019 Apr;231:56-62. doi: 10.1016/j.vetmic.2019.02.030. Epub 2019 Feb 22. Vet Microbiol. 2019. PMID: 30955824
-
Update on bacterial pathogenesis in BRD.Anim Health Res Rev. 2009 Dec;10(2):145-8. doi: 10.1017/S1466252309990193. Anim Health Res Rev. 2009. PMID: 20003651 Review.
-
Understanding the mechanisms of viral and bacterial coinfections in bovine respiratory disease: a comprehensive literature review of experimental evidence.Vet Res. 2022 Sep 6;53(1):70. doi: 10.1186/s13567-022-01086-1. Vet Res. 2022. PMID: 36068558 Free PMC article. Review.
Cited by
-
Sputum microbiota as a potential diagnostic marker for multidrug-resistant tuberculosis.Int J Med Sci. 2021 Mar 3;18(9):1935-1945. doi: 10.7150/ijms.53492. eCollection 2021. Int J Med Sci. 2021. PMID: 33850462 Free PMC article.
-
Transcript Characteristics on the Susceptibility Difference of Bovine Respiratory Disease.Int J Genomics. 2023 May 4;2023:9934684. doi: 10.1155/2023/9934684. eCollection 2023. Int J Genomics. 2023. PMID: 37180342 Free PMC article.
-
Effects of respiratory disease on Kele piglets lung microbiome, assessed through 16S rRNA sequencing.Vet World. 2020 Sep;13(9):1970-1981. doi: 10.14202/vetworld.2020.1970-1981. Epub 2020 Sep 25. Vet World. 2020. PMID: 33132613 Free PMC article.
-
Experimental challenge with bovine respiratory syncytial virus in dairy calves: bronchial lymph node transcriptome response.Sci Rep. 2019 Oct 14;9(1):14736. doi: 10.1038/s41598-019-51094-z. Sci Rep. 2019. PMID: 31611566 Free PMC article.
-
Updates and Current Challenges in Reproductive Microbiome: A Comparative Analysis between Cows and Women.Animals (Basel). 2024 Jul 3;14(13):1971. doi: 10.3390/ani14131971. Animals (Basel). 2024. PMID: 38998083 Free PMC article. Review.
References
-
- Brooks KR, Raper KC, Ward CE, Holland BP, Krehbiel CR, Step DL. Economic effects of bovine respiratory disease on feedlot cattle during backgrounding and finishing phases. Prof. Anim. Sci. 2011;27:195–203.
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials
Miscellaneous

