A case with concurrent duplication, triplication, and uniparental isodisomy at 1q42.12-qter supporting microhomology-mediated break-induced replication model for replicative rearrangements
- PMID: 28465723
- PMCID: PMC5410019
- DOI: 10.1186/s13039-017-0316-6
A case with concurrent duplication, triplication, and uniparental isodisomy at 1q42.12-qter supporting microhomology-mediated break-induced replication model for replicative rearrangements
Abstract
Background: Complex genomic rearrangements (CGRs) consisting of interstitial triplications in conjunction with uniparental isodisomy (isoUPD) have rarely been reported in patients with multiple congenital anomalies (MCA)/intellectual disability (ID). One-ended DNA break repair coupled with microhomology-mediated break-induced replication (MMBIR) has been recently proposed as a possible mechanism giving rise to interstitial copy number gains and distal isoUPD, although only a few cases providing supportive evidence in human congenital diseases with MCA have been documented.
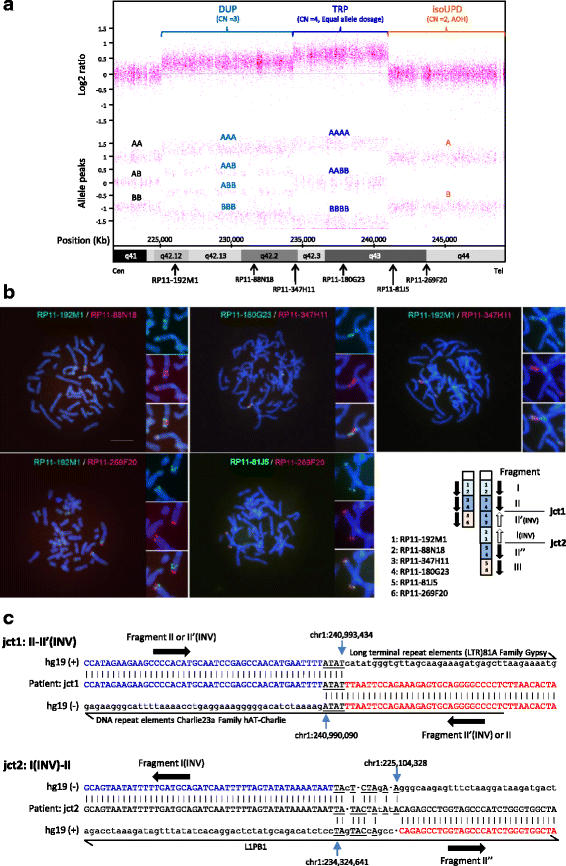
Case presentation: Here, we report on the chromosomal microarray (CMA)-based identification of the first known case with concurrent interstitial duplication at 1q42.12-q42.2 and triplication at 1q42.2-q43 followed by isoUPD for the remainder of chromosome 1q (at 1q43-qter). In distal 1q duplication/triplication overlapping with 1q42.12-q43, variable clinical features have been reported, and our 25-year-old patient with MCA/ID presented with some of these frequently described features. Further analyses including the precise mapping of breakpoint junctions within the CGR in a sequence level suggested that the CGR found in association with isoUPD in our case is a triplication with flanking duplications, characterized as a triplication with a particularly long duplication-inverted triplication-duplication (DUP-TRP/INV-DUP) structure. Because microhomology was observed in both junctions between the triplicated region and the flanking duplicated regions, our case provides supportive evidence for recently proposed replication-based mechanisms, such as MMBIR, underlying the formation of CGRs + isoUPD implicated in chromosomal disorders.
Conclusions: To the best of our knowledge, this is the first case of CGRs + isoUPD observed in 1q and having DUP-TRP/INV-DUP structure with a long proximal duplication, which supports MMBIR-based model for genomic rearrangements. Molecular cytogenetic analyses using CMA containing single-nucleotide polymorphism probes with further analyses of the breakpoint junctions are recommended in cases suspected of having complex chromosomal abnormalities based on discrepancies between clinical and conventional cytogenetic findings.
Keywords: 1q; Breakpoint junction sequence; Chromosomal microarray; Complex genomic rearrangement; DUP-TRP/INV-DUP structure; Microhomology-mediated break-induced replication model; Template switching; Uniparental isodisomy.
Figures

Similar articles
-
Concurrent triplication and uniparental isodisomy: evidence for microhomology-mediated break-induced replication model for genomic rearrangements.Eur J Hum Genet. 2015 Jan;23(1):61-6. doi: 10.1038/ejhg.2014.53. Epub 2014 Apr 9. Eur J Hum Genet. 2015. PMID: 24713661 Free PMC article.
-
Interchromosomal template-switching as a novel molecular mechanism for imprinting perturbations associated with Temple syndrome.Genome Med. 2019 Apr 23;11(1):25. doi: 10.1186/s13073-019-0633-y. Genome Med. 2019. PMID: 31014393 Free PMC article.
-
Complex genomic rearrangements at the PLP1 locus include triplication and quadruplication.PLoS Genet. 2015 Mar 6;11(3):e1005050. doi: 10.1371/journal.pgen.1005050. eCollection 2015 Mar. PLoS Genet. 2015. PMID: 25749076 Free PMC article.
-
Mechanisms of structural chromosomal rearrangement formation.Mol Cytogenet. 2022 Jun 14;15(1):23. doi: 10.1186/s13039-022-00600-6. Mol Cytogenet. 2022. PMID: 35701783 Free PMC article. Review.
-
A microhomology-mediated break-induced replication model for the origin of human copy number variation.PLoS Genet. 2009 Jan;5(1):e1000327. doi: 10.1371/journal.pgen.1000327. Epub 2009 Jan 30. PLoS Genet. 2009. PMID: 19180184 Free PMC article. Review.
Cited by
-
16p13.11p11.2 triplication syndrome: a new recognizable genomic disorder characterized by optical genome mapping and whole genome sequencing.Eur J Hum Genet. 2022 Jun;30(6):712-720. doi: 10.1038/s41431-022-01094-x. Epub 2022 Apr 7. Eur J Hum Genet. 2022. PMID: 35388186 Free PMC article.
-
Mechanisms and genomic implications of break-induced replication.Nat Struct Mol Biol. 2025 Aug 22. doi: 10.1038/s41594-025-01644-z. Online ahead of print. Nat Struct Mol Biol. 2025. PMID: 40846811 Review.
-
Triplication of the PCDH19 Gene as a Novel Disease Mechanism Leading to Epileptic Encephalopathy Resembling Loss-of-Function Pathogenic Variants.Genes (Basel). 2024 Oct 12;15(10):1312. doi: 10.3390/genes15101312. Genes (Basel). 2024. PMID: 39457436 Free PMC article.
-
A unifying model that explains the origins of human inverted copy number variants.PLoS Genet. 2024 Jan 4;20(1):e1011091. doi: 10.1371/journal.pgen.1011091. eCollection 2024 Jan. PLoS Genet. 2024. PMID: 38175827 Free PMC article. Review.
-
Clinical significance and mechanisms associated with segmental UPD.Mol Cytogenet. 2021 Jul 20;14(1):38. doi: 10.1186/s13039-021-00555-0. Mol Cytogenet. 2021. PMID: 34284807 Free PMC article.
References
-
- Beneteau C, Landais E, Doco-Fenzy M, Gavazzi C, Philippe C, Béri-Dexheimer M, et al. Microtriplication of 11q24.1: a highly recognisable phenotype with short stature, distinctive facial features, keratoconus, overweight, and intellectual disability. J Med Genet. 2011;48:635–9. doi: 10.1136/jmedgenet-2011-100008. - DOI - PubMed
Publication types
LinkOut - more resources
Full Text Sources
Other Literature Sources

