PRC2 represses transcribed genes on the imprinted inactive X chromosome in mice
- PMID: 28468635
- PMCID: PMC5415793
- DOI: 10.1186/s13059-017-1211-5
PRC2 represses transcribed genes on the imprinted inactive X chromosome in mice
Abstract
Background: Polycomb repressive complex 2 (PRC2) catalyzes histone H3K27me3, which marks many transcriptionally silent genes throughout the mammalian genome. Although H3K27me3 is associated with silenced gene expression broadly, it remains unclear why some but not other PRC2 target genes require PRC2 and H3K27me3 for silencing.
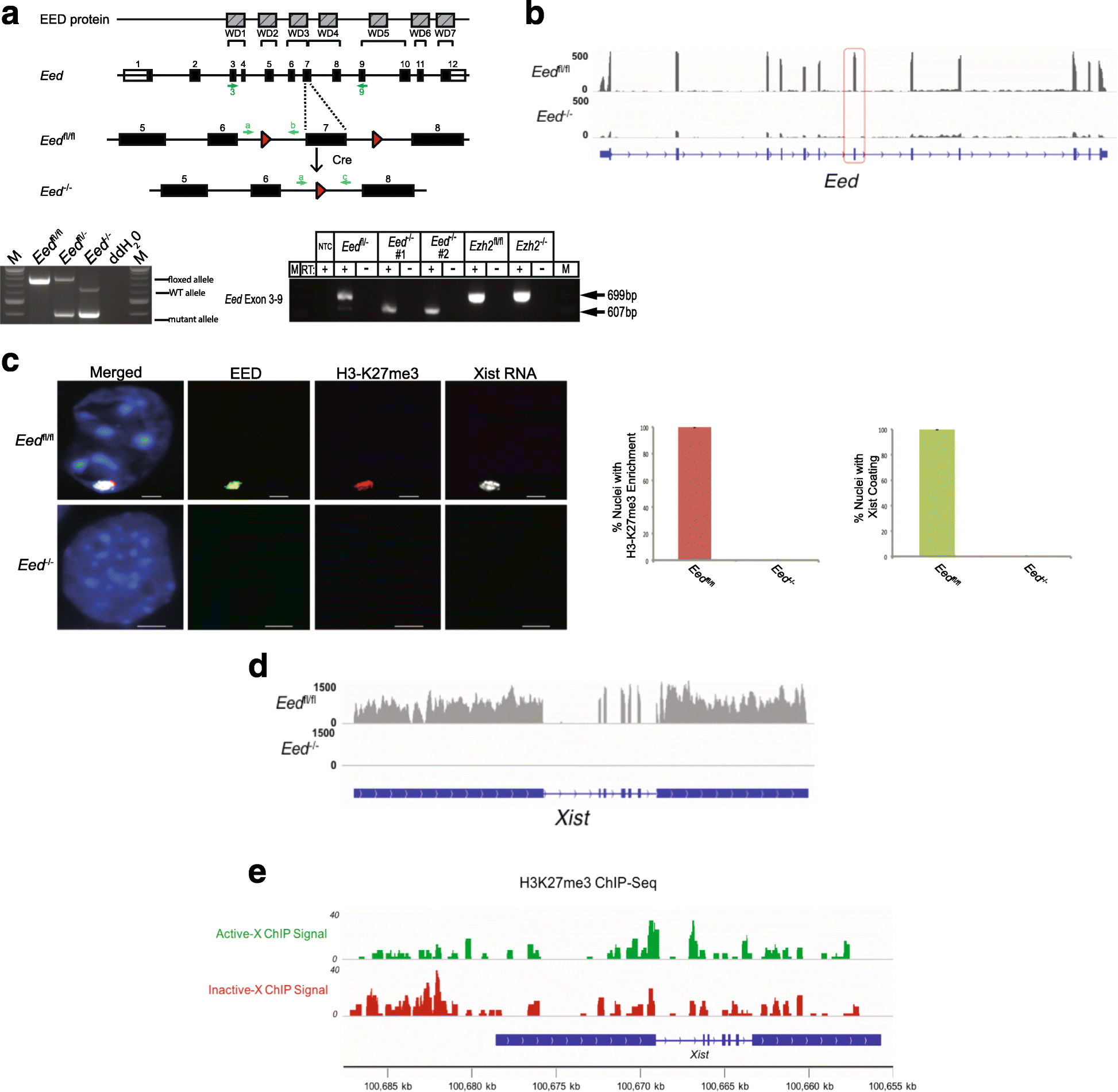
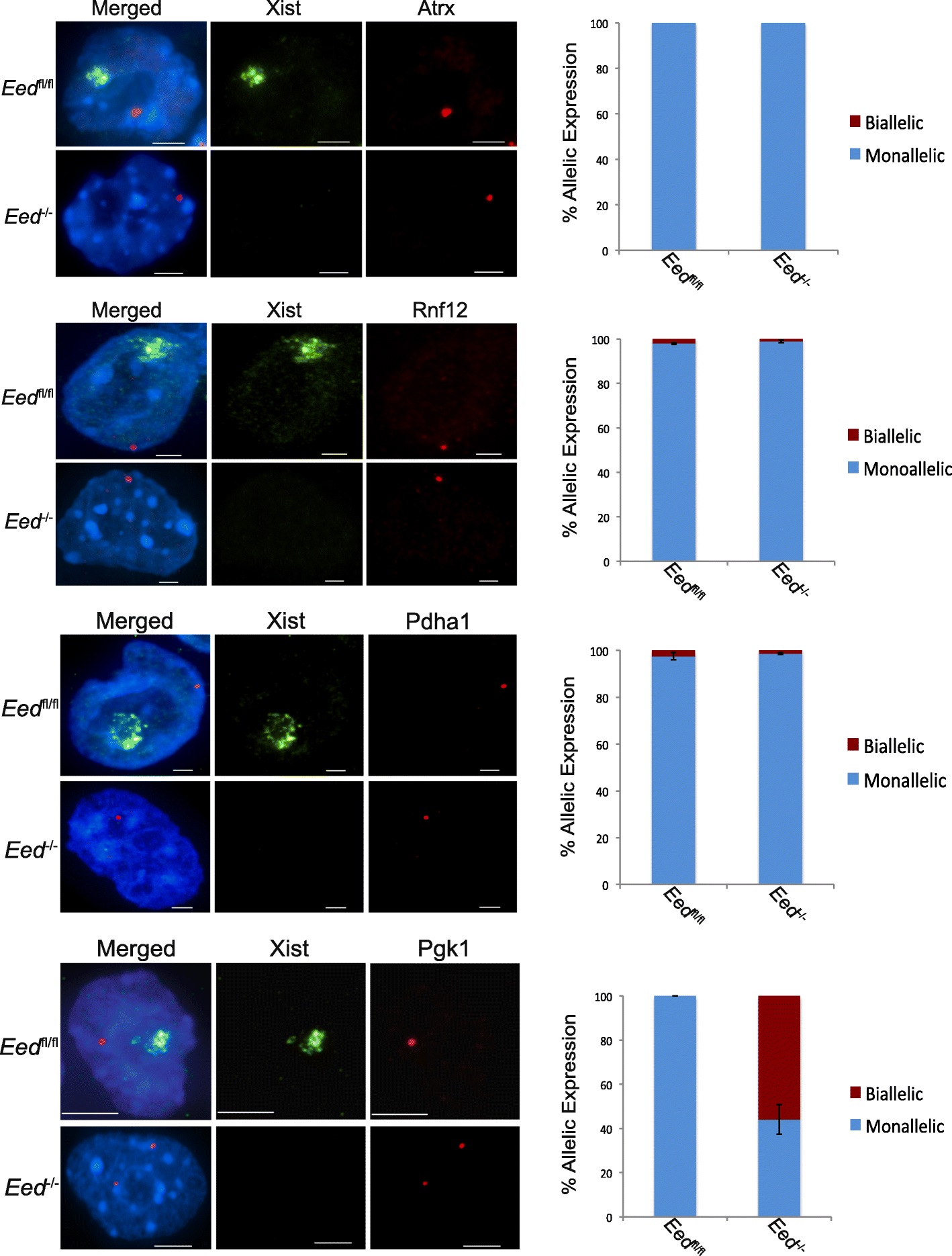
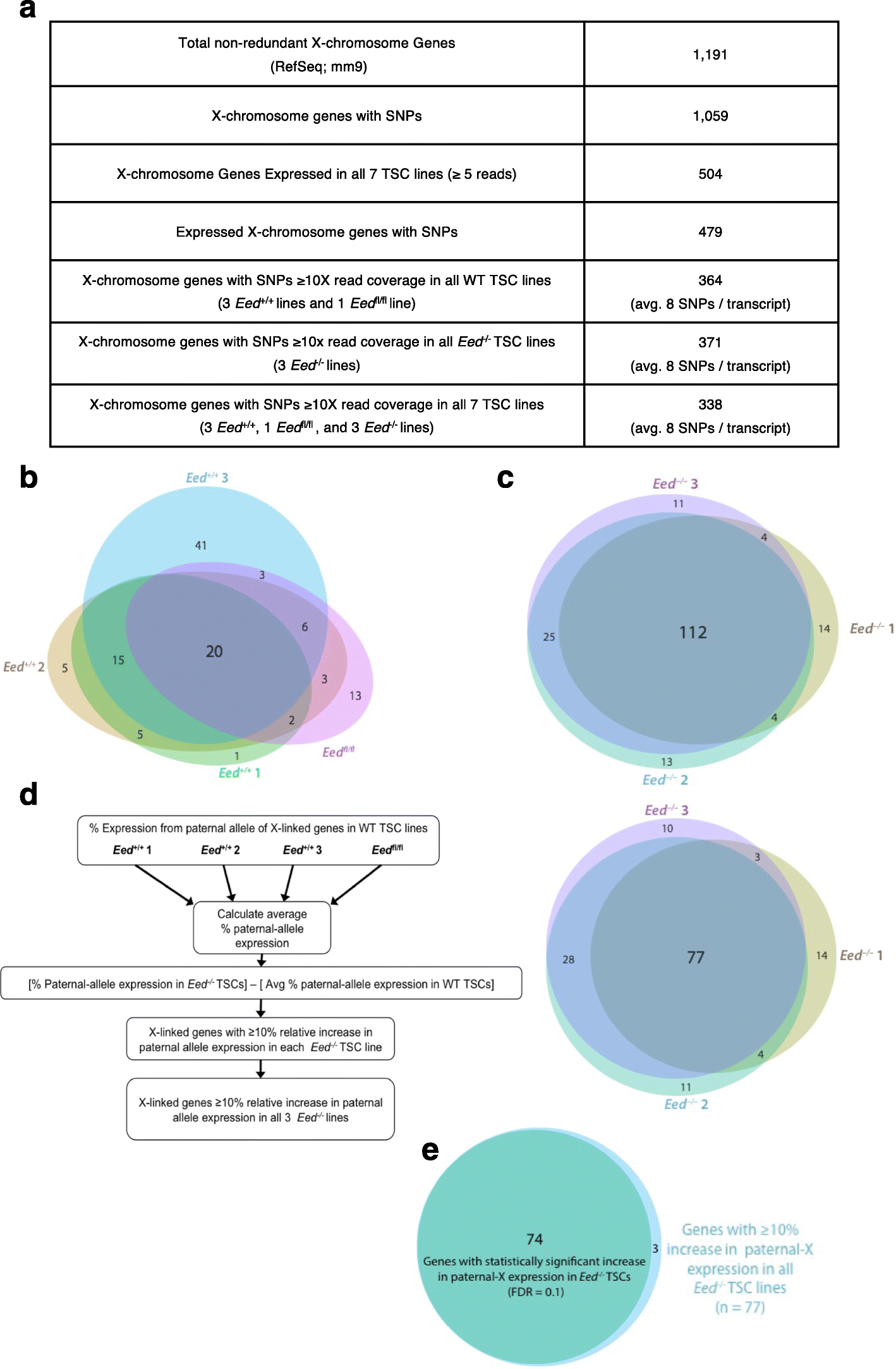
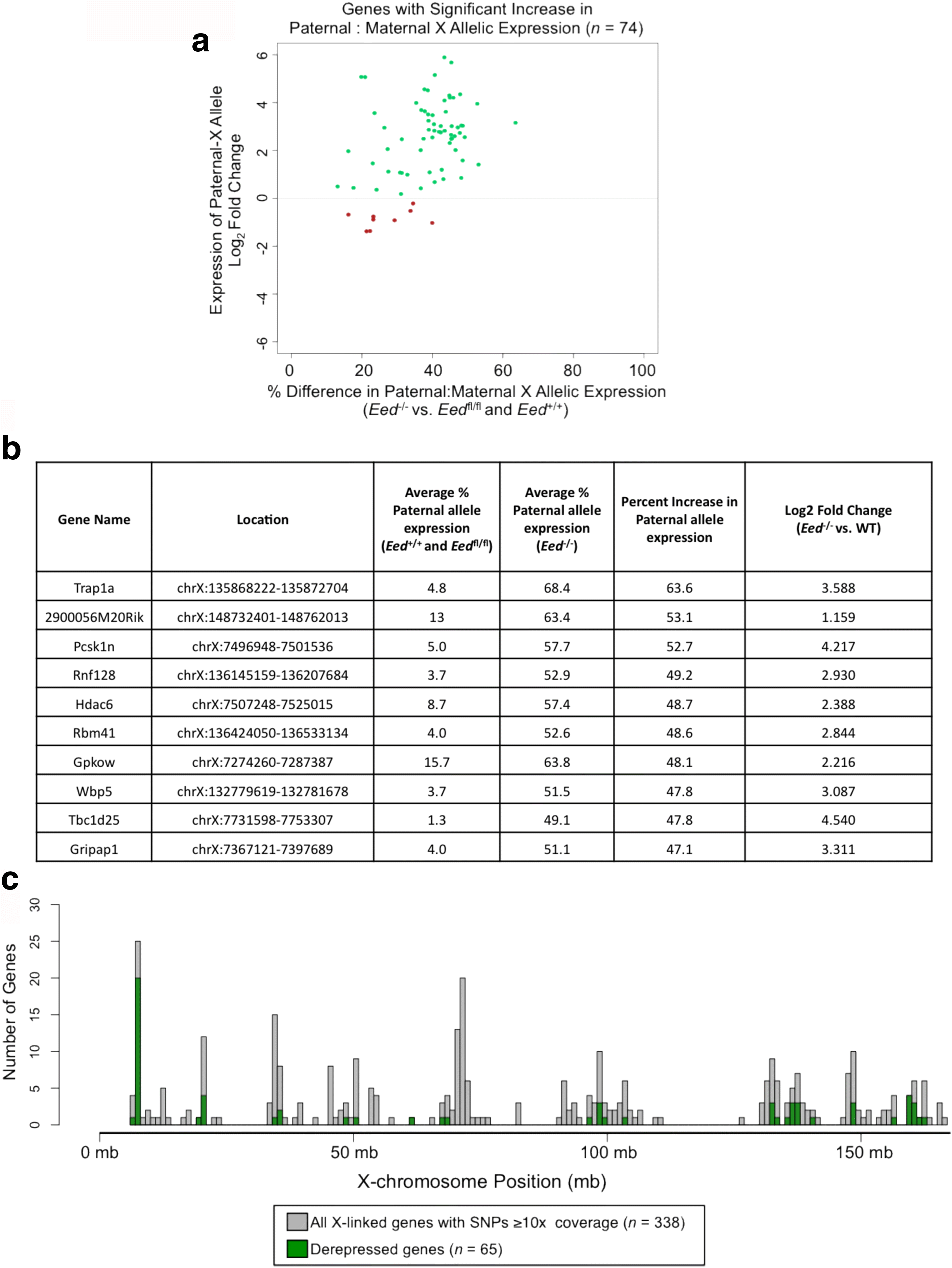
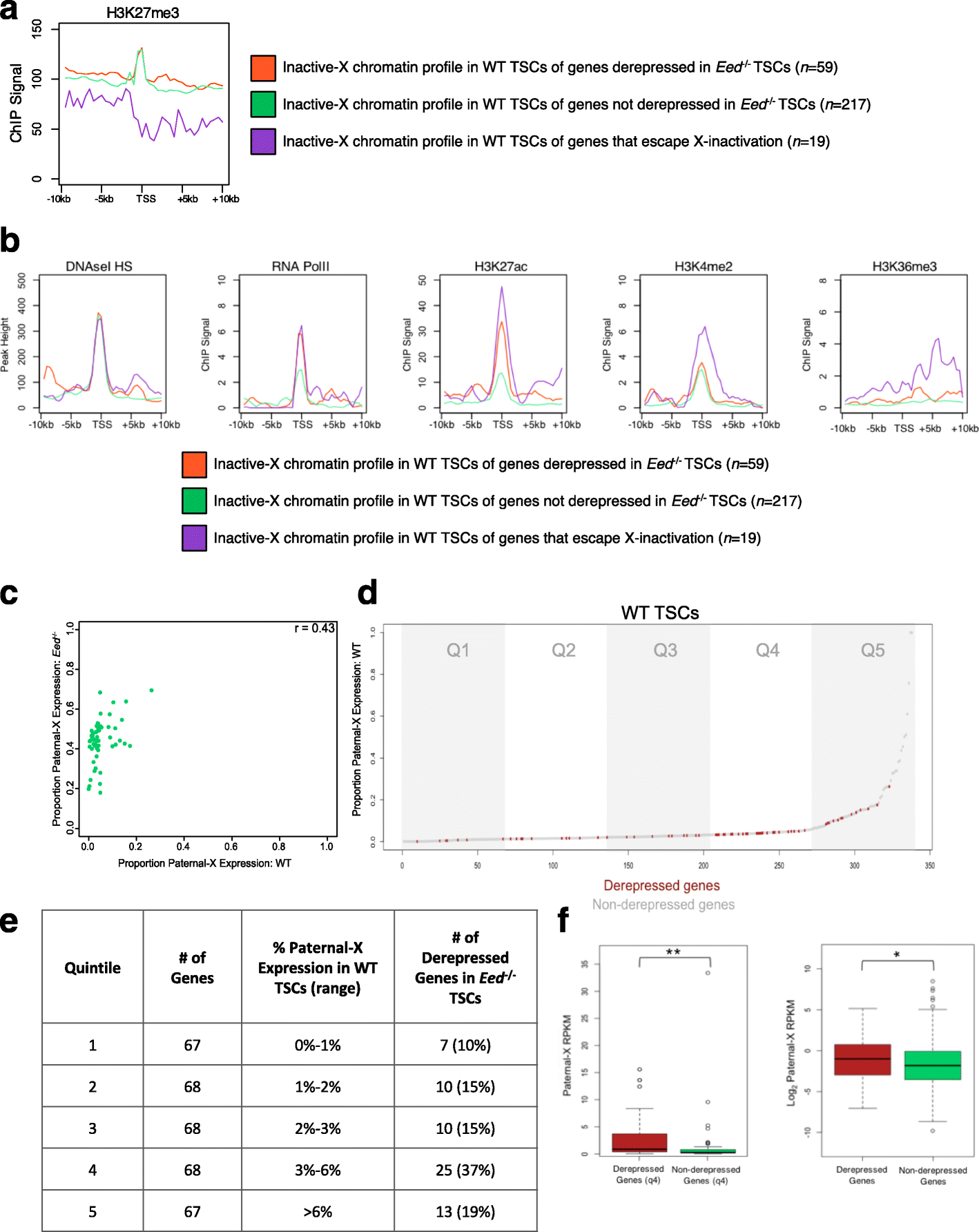
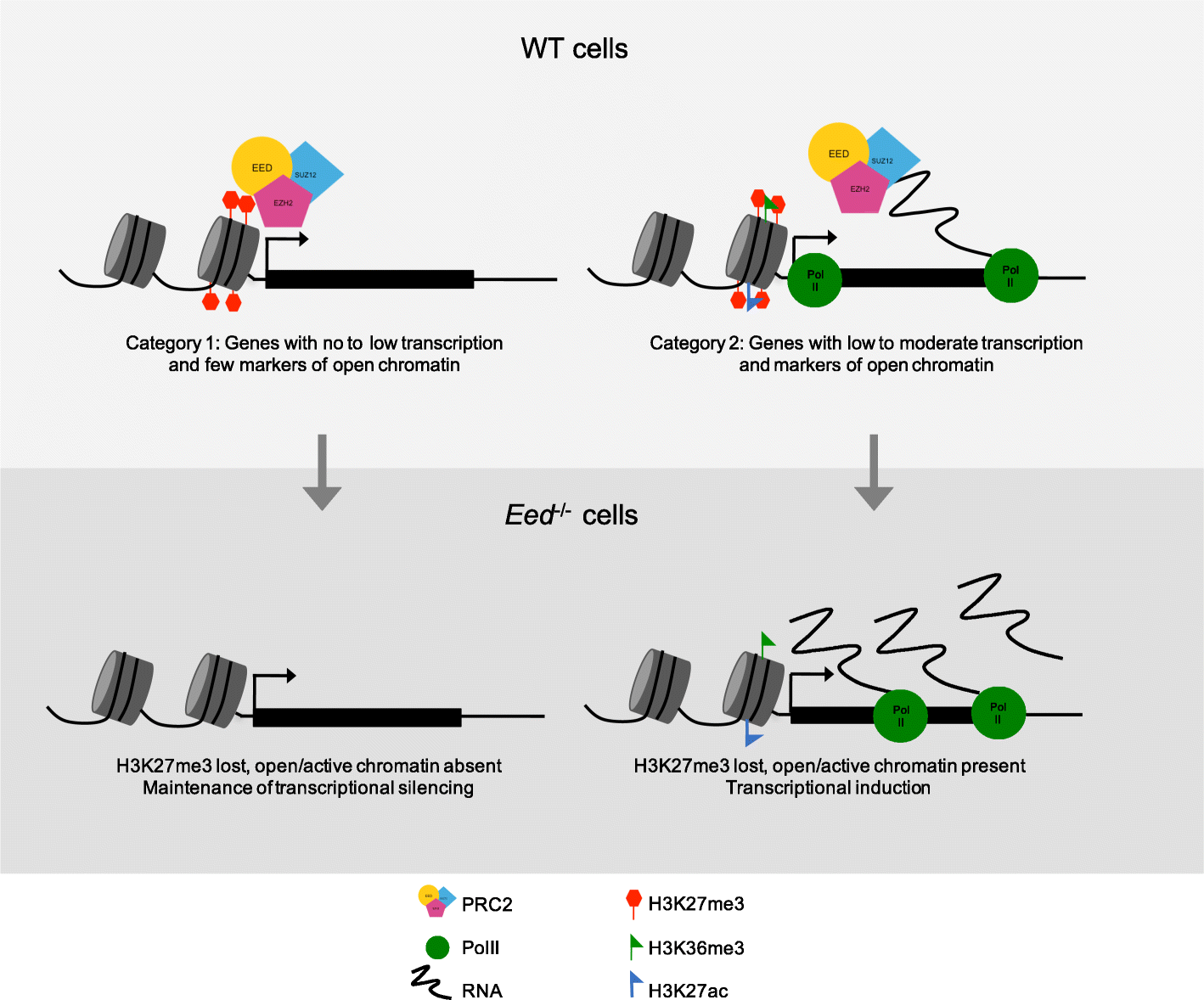
Results: Here we define the transcriptional and chromatin features that predict which PRC2 target genes require PRC2/H3K27me3 for silencing by interrogating imprinted mouse X-chromosome inactivation. H3K27me3 is enriched at promoters of silenced genes across the inactive X chromosome. To abrogate PRC2 function, we delete the core PRC2 protein EED in F1 hybrid trophoblast stem cells (TSCs), which undergo imprinted inactivation of the paternally inherited X chromosome. Eed -/- TSCs lack H3K27me3 and Xist lncRNA enrichment on the inactive X chromosome. Despite the absence of H3K27me3 and Xist RNA, only a subset of the inactivated X-linked genes is derepressed in Eed -/- TSCs. Unexpectedly, in wild-type (WT) TSCs these genes are transcribed and are enriched for active chromatin hallmarks on the inactive-X, including RNA PolII, H3K27ac, and H3K36me3, but not the bivalent mark H3K4me2. By contrast, PRC2 targets that remain repressed in Eed -/- TSCs are depleted for active chromatin characteristics in WT TSCs.
Conclusions: A comparative analysis of transcriptional and chromatin features of inactive X-linked genes in WT and Eed -/- TSCs suggests that PRC2 acts as a brake to prevent induction of transcribed genes on the inactive X chromosome, a mode of PRC2 function that may apply broadly.
Keywords: EED; H3K27me3; Imprinting; PRC2; Polycomb; Trophoblast stem cells; X-inactivation.
Figures
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

