Transposable Elements in Human Cancer: Causes and Consequences of Deregulation
- PMID: 28471386
- PMCID: PMC5454887
- DOI: 10.3390/ijms18050974
Transposable Elements in Human Cancer: Causes and Consequences of Deregulation
Abstract
Transposable elements (TEs) comprise nearly half of the human genome and play an essential role in the maintenance of genomic stability, chromosomal architecture, and transcriptional regulation. TEs are repetitive sequences consisting of RNA transposons, DNA transposons, and endogenous retroviruses that can invade the human genome with a substantial contribution in human evolution and genomic diversity. TEs are therefore firmly regulated from early embryonic development and during the entire course of human life by epigenetic mechanisms, in particular DNA methylation and histone modifications. The deregulation of TEs has been reported in some developmental diseases, as well as for different types of human cancers. To date, the role of TEs, the mechanisms underlying TE reactivation, and the interplay with DNA methylation in human cancers remain largely unexplained. We reviewed the loss of epigenetic regulation and subsequent genomic instability, chromosomal aberrations, transcriptional deregulation, oncogenic activation, and aberrations of non-coding RNAs as the potential mechanisms underlying TE deregulation in human cancers.
Keywords: cancer; epigenetics; genomic instability; non-coding RNAs; transposable elements.
Conflict of interest statement
The authors declare no conflict of interest.
Figures

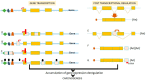
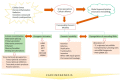
References
-
- Lu X.-J., Xue H.-Y., Qi X., Xu J., Ma S.-J. LINE-1 in cancer: Multifaceted functions and potential clinical implications. Genet. Med. 2016;18:431–439. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

