A mechanistic study of helicases with magnetic traps
- PMID: 28474797
- PMCID: PMC5477542
- DOI: 10.1002/pro.3187
A mechanistic study of helicases with magnetic traps
Abstract
Helicases are a broad family of enzymes that separate nucleic acid double strand structures (DNA/DNA, DNA/RNA, or RNA/RNA) and thus are essential to DNA replication and the maintenance of nucleic acid integrity. We review the picture that has emerged from single molecule studies of the mechanisms of DNA and RNA helicases and their interactions with other proteins. Many features have been uncovered by these studies that were obscured by bulk studies, such as DNA strands switching, mechanical (rather than biochemical) coupling between helicases and polymerases, helicase-induced re-hybridization and stalled fork rescue.
Keywords: DNA unwinding; Holliday junction migration; active unwinding; ds-DNA fork; fork regression; helicase/polymerase coupling; helicases; magnetic traps; polymerase; primase; primosome; replisome; strand annealing.
© 2017 The Protein Society.
Figures





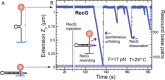
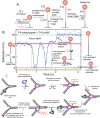

References
-
- Delagoutte E, von Hippel PH (2002) Helicase mechanisms and the coupling of helicases within macromolecular machines Part 1: Structures and properties of isolated helicases. Quart Rev Biophys 35:431–478. - PubMed
-
- Delagoutte E, von Hippel PH (2003) Helicase mechanisms and the coupling of helicases within macromolecular machines ‐ Part II: Integration of helicases into cellular processes. Quart Rev Biophys 36:1–69. - PubMed
-
- Lohman TM, Bjornson KP (1996) Mechanisms of helicase‐catalyzed unwinding. Annu Rev Biochem 65:169–214. - PubMed
-
- West SC (1996) DNA helicases: New breeds of translocating motors and molecular pumps. Cell 86:177–180. - PubMed
-
- Soultanas P, Wigley DB (2001) Unwinding the ‘Gordian knot’ of helicase action. Trends Biochem Sci 26:47–54. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

