Artemether-Lumefantrine and Dihydroartemisinin-Piperaquine Exert Inverse Selective Pressure on Plasmodium Falciparum Drug Sensitivity-Associated Haplotypes in Uganda
- PMID: 28480232
- PMCID: PMC5413987
- DOI: 10.1093/ofid/ofw229
Artemether-Lumefantrine and Dihydroartemisinin-Piperaquine Exert Inverse Selective Pressure on Plasmodium Falciparum Drug Sensitivity-Associated Haplotypes in Uganda
Abstract
Background: Altered sensitivity to multiple antimalarial drugs is mediated by polymorphisms in pfmdr1, which encodes the Plasmodium falciparum multidrug resistance transporter. In Africa the N86Y and D1246Y polymorphisms have been shown to be selected by treatment, with artemether-lumefantrine (AL) and dihydroartemisinin-piperaquine (DP) selecting for wild-type and mutant alleles, respectively. However, there has been little study of pfmdr1 haplotypes, in part because haplotype analyses are complicated by multiclonal infections.
Methods: We fit a haplotype frequency estimation model, which accounts for multiclonal infections, to the polymorphic pfmdr1 N86Y, Y184F, and D1246Y alleles in samples from a longitudinal trial comparing AL and DP to treat uncomplicated P falciparum malaria in Tororo, Uganda from 2007 to 2012. We regressed estimates onto covariates of trial arm and selective drug pressure.
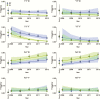
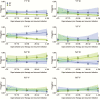
Results: Yearly trends showed increasing frequency estimates for haplotypes with wild type pfmdr1 N86 and D1246 alleles and decreasing frequency estimates for haplotypes with the mutant pfmdr1 86Y allele. Considering days since prior therapy, we saw evidence suggestive of selection by AL for haplotypes with N86 combined with 184F, D1246, or both, and against all haplotypes with 86Y, and evidence suggestive of selection by DP for 86Y only when combined with Y184 and 1246Y (haplotype YYY) and against haplotypes NFD and NYY.
Conclusions: Based on our model, AL selected several haplotypes containing N86, whereas DP selection was haplotype specific, demonstrating the importance of haplotype analyses. Inverse selective pressure of AL and DP on the complementary haplotypes NFD and YYY suggests that rotating artemisinin-based antimalarial combination regimens may be the best treatment option to prevent resistance selection.
Keywords: Uganda.; artemether-lumefantrine; dihydroartemisinin-piperaquine; drug resistance; haplotype.
Figures
References
-
- World Health Organization. World Malaria Report 2015. Geneva: World Health Organization; 2015.
-
- World Health Organization. Guidelines for the Treatment of Malaria. 3rd edition Geneva: World Health Organization; 2015.
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

