Hepatic p63 regulates steatosis via IKKβ/ER stress
- PMID: 28480888
- PMCID: PMC5424198
- DOI: 10.1038/ncomms15111
Hepatic p63 regulates steatosis via IKKβ/ER stress
Erratum in
-
Corrigendum: Hepatic p63 regulates steatosis via IKKβ/ER stress.Nat Commun. 2017 Jun 16;8:16059. doi: 10.1038/ncomms16059. Nat Commun. 2017. PMID: 28621331 Free PMC article.
Abstract
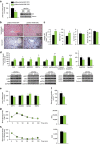
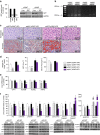
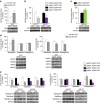
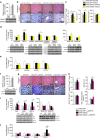
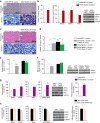
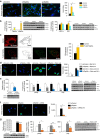
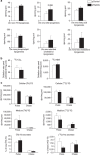
p53 family members control several metabolic and cellular functions. The p53 ortholog p63 modulates cellular adaptations to stress and has a major role in cell maintenance and proliferation. Here we show that p63 regulates hepatic lipid metabolism. Mice with liver-specific p53 deletion develop steatosis and show increased levels of p63. Down-regulation of p63 attenuates liver steatosis in p53 knockout mice and in diet-induced obese mice, whereas the activation of p63 induces lipid accumulation. Hepatic overexpression of N-terminal transactivation domain TAp63 induces liver steatosis through IKKβ activation and the induction of ER stress, the inhibition of which rescues the liver functions. Expression of TAp63, IKKβ and XBP1s is also increased in livers of obese patients with NAFLD. In cultured human hepatocytes, TAp63 inhibition protects against oleic acid-induced lipid accumulation, whereas TAp63 overexpression promotes lipid storage, an effect reversible by IKKβ silencing. Our findings indicate an unexpected role of the p63/IKKβ/ER stress pathway in lipid metabolism and liver disease.
Conflict of interest statement
The authors declare no competing financial interests.
Figures



References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials
Miscellaneous

