The Tetrazole VT-1161 Is a Potent Inhibitor of Trichophyton rubrum through Its Inhibition of T. rubrum CYP51
- PMID: 28483956
- PMCID: PMC5487648
- DOI: 10.1128/AAC.00333-17
The Tetrazole VT-1161 Is a Potent Inhibitor of Trichophyton rubrum through Its Inhibition of T. rubrum CYP51
Abstract
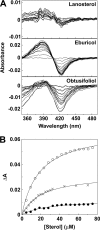
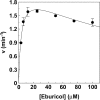
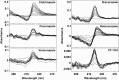
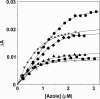
Prior to characterization of antifungal inhibitors that target CYP51, Trichophyton rubrum CYP51 was expressed in Escherichia coli, purified, and characterized. T. rubrum CYP51 bound lanosterol, obtusifoliol, and eburicol with similar affinities (dissociation constant [Kd ] values, 22.7, 20.3, and 20.9 μM, respectively) but displayed substrate specificity, insofar as only eburicol was demethylated in CYP51 reconstitution assays (turnover number, 1.55 min-1; Km value, 2 μM). The investigational agent VT-1161 bound tightly to T. rubrum CYP51 (Kd = 242 nM) with an affinity similar to that of clotrimazole, fluconazole, ketoconazole, and voriconazole (Kd values, 179, 173, 312, and 304 nM, respectively) and with an affinity lower than that of itraconazole (Kd = 53 nM). Determinations of 50% inhibitory concentrations (IC50s) using 0.5 μM CYP51 showed that VT-1161 was a tight-binding inhibitor of T. rubrum CYP51 activity, yielding an IC50 of 0.14 μM, whereas itraconazole, fluconazole, and ketoconazole had IC50s of 0.26, 0.4, and 0.6 μM, respectively. When the activity of VT-1161 was tested against 34 clinical isolates, VT-1161 was a potent inhibitor of T. rubrum growth, with MIC50, MIC90, and geometric mean MIC values of ≤0.03, 0.06, and 0.033 μg ml-1, respectively. With its selectivity versus human CYP51 and drug-metabolizing cytochrome P450s having already been established, VT-1161 should prove to be safe and effective in combating T. rubrum infections in patients.
Keywords: CYP51; Trichophyton rubrum; VT-1161; azole resistance; substrate specificity.
Copyright © 2017 American Society for Microbiology.
Figures


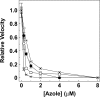
References
-
- Summerbell RC, Kane J, Krajden S. 1989. Onychomycosis, tinea pedis and tinea manuum caused by non-dermatophytic filamentous fungi. Mycoses 32:609–619. - PubMed
-
- Tietz HJ, Kunzelmann V, Schonian G. 1995. Changes in the fungal spectrum of dermatomycoses. Mycoses 38(Suppl 1):S33–S39. - PubMed
-
- Santos DA, Hamdan JS. 2006. In vitro antifungal oral drug and drug-combination activity against onychomycosis causative dermatophytes. Med Mycol 44:357–362. - PubMed
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

