Characterization of the Transcriptomes of Lgr5+ Hair Cell Progenitors and Lgr5- Supporting Cells in the Mouse Cochlea
- PMID: 28491023
- PMCID: PMC5405134
- DOI: 10.3389/fnmol.2017.00122
Characterization of the Transcriptomes of Lgr5+ Hair Cell Progenitors and Lgr5- Supporting Cells in the Mouse Cochlea
Abstract
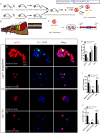
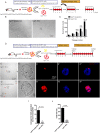
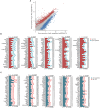
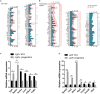
Cochlear supporting cells (SCs) have been shown to be a promising resource for hair cell (HC) regeneration in the neonatal mouse cochlea. Previous studies have reported that Lgr5+ SCs can regenerate HCs both in vitro and in vivo and thus are considered to be inner ear progenitor cells. Lgr5+ progenitors are able to regenerate more HCs than Lgr5- SCs, and it is important to understand the mechanism behind the proliferation and HC regeneration of these progenitors. Here, we isolated Lgr5+ progenitors and Lgr5- SCs from Lgr5-EGFP-CreERT2/Sox2-CreERT2/Rosa26-tdTomato mice via flow cytometry. As expected, we found that Lgr5+ progenitors had significantly higher proliferation and HC regeneration ability than Lgr5- SCs. Next, we performed RNA-Seq to determine the gene expression profiles of Lgr5+ progenitors and Lgr5- SCs. We analyzed the genes that were enriched and differentially expressed in Lgr5+ progenitors and Lgr5- SCs, and we found 8 cell cycle genes, 9 transcription factors, and 24 cell signaling pathway genes that were uniquely expressed in one population but not the other. Last, we made a protein-protein interaction network to further analyze the role of these differentially expressed genes. In conclusion, we present a set of genes that might regulate the proliferation and HC regeneration ability of Lgr5+ progenitors, and these might serve as potential new therapeutic targets for HC regeneration.
Keywords: RNA-Seq; differentiation; gene expression; proliferation; regeneration; sphere.
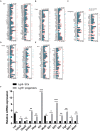
Figures


Similar articles
-
Characterization of Lgr5+ Progenitor Cell Transcriptomes after Neomycin Injury in the Neonatal Mouse Cochlea.Front Mol Neurosci. 2017 Jul 4;10:213. doi: 10.3389/fnmol.2017.00213. eCollection 2017. Front Mol Neurosci. 2017. PMID: 28725177 Free PMC article.
-
Characterization of Lgr5+ progenitor cell transcriptomes in the apical and basal turns of the mouse cochlea.Oncotarget. 2016 Jul 5;7(27):41123-41141. doi: 10.18632/oncotarget.8636. Oncotarget. 2016. PMID: 27070092 Free PMC article.
-
Age-related transcriptome changes in Sox2+ supporting cells in the mouse cochlea.Stem Cell Res Ther. 2019 Dec 2;10(1):365. doi: 10.1186/s13287-019-1437-0. Stem Cell Res Ther. 2019. PMID: 31791390 Free PMC article.
-
Hair cell regeneration from inner ear progenitors in the mammalian cochlea.Am J Stem Cells. 2020 Jun 15;9(3):25-35. eCollection 2020. Am J Stem Cells. 2020. PMID: 32699655 Free PMC article. Review.
-
Understanding the differentiation and epigenetics of cochlear sensory progenitors in pursuit of regeneration.Curr Opin Otolaryngol Head Neck Surg. 2021 Oct 1;29(5):366-372. doi: 10.1097/MOO.0000000000000741. Curr Opin Otolaryngol Head Neck Surg. 2021. PMID: 34374667 Free PMC article. Review.
Cited by
-
The differentiation of Lgr5+ progenitor cells on nanostructures of self-assembled silica beads.PLoS One. 2024 Jul 12;19(7):e0304809. doi: 10.1371/journal.pone.0304809. eCollection 2024. PLoS One. 2024. PMID: 38995923 Free PMC article.
-
Transcriptome sequencing of adenomyosis eutopic endometrium: A new insight into its pathophysiology.J Cell Mol Med. 2019 Dec;23(12):8381-8391. doi: 10.1111/jcmm.14718. Epub 2019 Oct 1. J Cell Mol Med. 2019. PMID: 31576674 Free PMC article.
-
Hollow Mesoporous Silica@Zeolitic Imidazolate Framework Capsules and Their Applications for Gentamicin Delivery.Neural Plast. 2018 Apr 5;2018:2160854. doi: 10.1155/2018/2160854. eCollection 2018. Neural Plast. 2018. PMID: 29849553 Free PMC article.
-
Characterization of Lgr6+ Cells as an Enriched Population of Hair Cell Progenitors Compared to Lgr5+ Cells for Hair Cell Generation in the Neonatal Mouse Cochlea.Front Mol Neurosci. 2018 May 14;11:147. doi: 10.3389/fnmol.2018.00147. eCollection 2018. Front Mol Neurosci. 2018. PMID: 29867341 Free PMC article.
-
Junctional E-cadherin/p120-catenin Is Correlated with the Absence of Supporting Cells to Hair Cells Conversion in Postnatal Mice Cochleae.Front Mol Neurosci. 2018 Feb 21;11:20. doi: 10.3389/fnmol.2018.00020. eCollection 2018. Front Mol Neurosci. 2018. PMID: 29515364 Free PMC article.
References
-
- Alvarado D. M., Hawkins R. D., Bashiardes S., Veile R. A., Ku Y. C., Powder K. E., et al. (2011). An RNA interference-based screen of transcription factor genes identifies pathways necessary for sensory regeneration in the avian inner ear. J. Neurosci. 31 4535–4543. 10.1523/JNEUROSCI.5456-10.2011 - DOI - PMC - PubMed
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials

