Biomolecular engineering for nanobio/bionanotechnology
- PMID: 28491487
- PMCID: PMC5401866
- DOI: 10.1186/s40580-017-0103-4
Biomolecular engineering for nanobio/bionanotechnology
Abstract
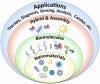
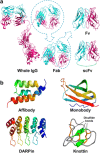
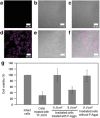
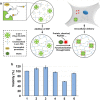
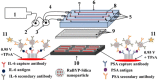
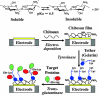
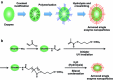
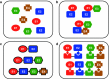
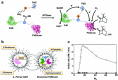
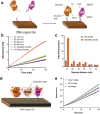
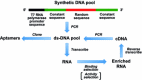
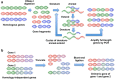
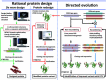
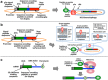
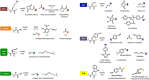
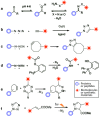
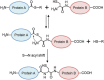
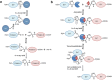
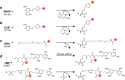
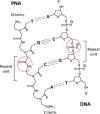
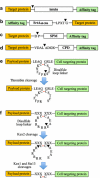
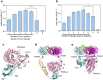
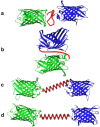
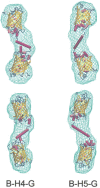
Biomolecular engineering can be used to purposefully manipulate biomolecules, such as peptides, proteins, nucleic acids and lipids, within the framework of the relations among their structures, functions and properties, as well as their applicability to such areas as developing novel biomaterials, biosensing, bioimaging, and clinical diagnostics and therapeutics. Nanotechnology can also be used to design and tune the sizes, shapes, properties and functionality of nanomaterials. As such, there are considerable overlaps between nanotechnology and biomolecular engineering, in that both are concerned with the structure and behavior of materials on the nanometer scale or smaller. Therefore, in combination with nanotechnology, biomolecular engineering is expected to open up new fields of nanobio/bionanotechnology and to contribute to the development of novel nanobiomaterials, nanobiodevices and nanobiosystems. This review highlights recent studies using engineered biological molecules (e.g., oligonucleotides, peptides, proteins, enzymes, polysaccharides, lipids, biological cofactors and ligands) combined with functional nanomaterials in nanobio/bionanotechnology applications, including therapeutics, diagnostics, biosensing, bioanalysis and biocatalysts. Furthermore, this review focuses on five areas of recent advances in biomolecular engineering: (a) nucleic acid engineering, (b) gene engineering, (c) protein engineering, (d) chemical and enzymatic conjugation technologies, and (e) linker engineering. Precisely engineered nanobiomaterials, nanobiodevices and nanobiosystems are anticipated to emerge as next-generation platforms for bioelectronics, biosensors, biocatalysts, molecular imaging modalities, biological actuators, and biomedical applications.
Keywords: Bioanalysis; Biocatalyst; Biosensing; Conjugation technologies; Diagnosis; Engineered biological molecules; Gene engineering; Nucleic acid engineering; Protein engineering; Therapy.
Figures





Similar articles
-
Recent progress in biomolecular engineering.Biotechnol Prog. 2000 Jan-Feb;16(1):2-16. doi: 10.1021/bp088059d. Biotechnol Prog. 2000. PMID: 10662483 Review.
-
Construction of rolling circle amplification products-based pure nucleic acid nanostructures for biomedical applications.Acta Biomater. 2023 Apr 1;160:1-13. doi: 10.1016/j.actbio.2023.02.005. Epub 2023 Feb 9. Acta Biomater. 2023. PMID: 36764595 Review.
-
Atomic force microscopy as an imaging tool to study the bio/nonbio complexes.J Microsc. 2020 Dec;280(3):241-251. doi: 10.1111/jmi.12936. Epub 2020 Jun 17. J Microsc. 2020. PMID: 32519330
-
Engineering DNA on the Surface of Upconversion Nanoparticles for Bioanalysis and Therapeutics.ACS Nano. 2021 Nov 23;15(11):17257-17274. doi: 10.1021/acsnano.1c08036. Epub 2021 Nov 12. ACS Nano. 2021. PMID: 34766752 Review.
-
Nanobio interfaces: charge control of enzyme/inorganic interfaces for advanced biocatalysis.Langmuir. 2013 Nov 19;29(46):14001-16. doi: 10.1021/la403165y. Epub 2013 Oct 24. Langmuir. 2013. PMID: 24102555 Review.
Cited by
-
Discrimination and isolation of the virus from free RNA fragments for the highly sensitive measurement of SARS-CoV-2 abundance on surfaces using a graphene oxide nano surface.Nano Converg. 2021 Oct 18;8(1):31. doi: 10.1186/s40580-021-00281-8. Nano Converg. 2021. PMID: 34661769 Free PMC article.
-
Impact of the conjugation of antibodies to the surfaces of polymer nanoparticles on the immune cell targeting abilities.Nano Converg. 2021 Aug 16;8(1):24. doi: 10.1186/s40580-021-00274-7. Nano Converg. 2021. PMID: 34398322 Free PMC article.
-
Progress and Prospects of Biomolecular Materials in Solar Photovoltaic Applications.Molecules. 2025 Aug 1;30(15):3236. doi: 10.3390/molecules30153236. Molecules. 2025. PMID: 40807408 Free PMC article. Review.
-
Analysis of DNA Origami Nanostructures Using Capillary Electrophoresis.Anal Chem. 2023 Dec 26;95(51):18783-18792. doi: 10.1021/acs.analchem.3c03641. Epub 2023 Dec 13. Anal Chem. 2023. PMID: 38088564 Free PMC article.
-
CD19 Chimeric Antigen Receptor-Exosome Targets CD19 Positive B-lineage Acute Lymphocytic Leukemia and Induces Cytotoxicity.Cancers (Basel). 2021 Mar 19;13(6):1401. doi: 10.3390/cancers13061401. Cancers (Basel). 2021. PMID: 33808645 Free PMC article.
References
Publication types
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials
